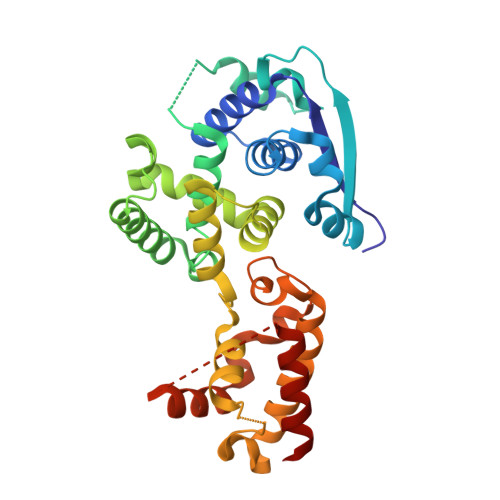
Insight into the Ebola virus nucleocapsid assembly mechanism: crystal structure of Ebola virus nucleoprotein core domain at 1.8 A resolution.
Dong, S., Yang, P., Li, G., Liu, B., Wang, W., Liu, X., Xia, B., Yang, C., Lou, Z., Guo, Y., Rao, Z.(2015) Protein Cell 6: 351-362
- PubMed: 25910597
- DOI: https://doi.org/10.1007/s13238-015-0163-3
- Primary Citation of Related Structures:
4Z9P - PubMed Abstract:
Ebola virus (EBOV) is a key member of Filoviridae family and causes severe human infectious diseases with high morbidity and mortality. As a typical negative-sense single-stranded RNA (-ssRNA) viruses, EBOV possess a nucleocapsid protein (NP) to facilitate genomic RNA encapsidation to form viral ribonucleoprotein complex (RNP) together with genome RNA and polymerase, which plays the most essential role in virus proliferation cycle. However, the mechanism of EBOV RNP formation remains unclear. In this work, we solved the high resolution structure of core domain of EBOV NP. The polypeptide of EBOV NP core domain (NP(core)) possesses an N-lobe and C-lobe to clamp a RNA binding groove, presenting similarities with the structures of the other reported viral NPs encoded by the members from Mononegavirales order. Most strikingly, a hydrophobic pocket at the surface of the C-lobe is occupied by an α-helix of EBOV NP(core) itself, which is highly conserved among filoviridae family. Combined with other biochemical and biophysical evidences, our results provides great potential for understanding the mechanism underlying EBOV RNP formation via the mobility of EBOV NP element and enables the development of antiviral therapies targeting EBOV RNP formation.
Organizational Affiliation:
State Key Laboratory of Medicinal Chemical Biology and College of Pharmacy, Nankai University, Tianjin, 300071, China.