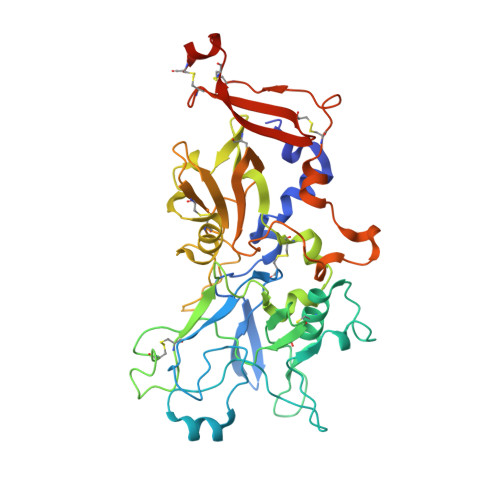
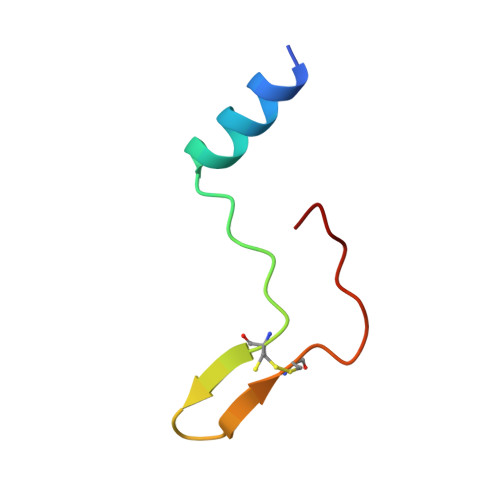
An Extended Surface Loop on Toxoplasma gondii Apical Membrane Antigen 1 (AMA1) Governs Ligand Binding Selectivity.
Parker, M.L., Boulanger, M.J.(2015) PLoS One 10: e0126206-e0126206
- PubMed: 25955165
- DOI: https://doi.org/10.1371/journal.pone.0126206
- Primary Citation of Related Structures:
4YIV, 4YIZ - PubMed Abstract:
Apicomplexan parasites are the causative agents of globally prevalent diseases including malaria and toxoplasmosis. These obligate intracellular pathogens have evolved a sophisticated host cell invasion strategy that relies on a parasite-host cell junction anchored by interactions between apical membrane antigens (AMAs) on the parasite surface and rhoptry neck 2 (RON2) proteins discharged from the parasite and embedded in the host cell membrane. Key to formation of the AMA1-RON2 complex is displacement of an extended surface loop on AMA1 called the DII loop. While conformational flexibility of the DII loop is required to expose the mature RON2 binding groove, a definitive role of this substructure has not been elucidated. To establish a role of the DII loop in Toxoplasma gondii AMA1, we engineered a form of the protein where the mobile portion of the loop was replaced with a short Gly-Ser linker (TgAMA1ΔDIIloop). Isothermal titration calorimetry measurements with a panel of RON2 peptides revealed an influential role for the DII loop in governing selectivity. Most notably, an Eimeria tenella RON2 (EtRON2) peptide that showed only weak binding to TgAMA1 bound with high affinity to TgAMA1ΔDIIloop. To define the molecular basis for the differential binding, we determined the crystal structure of TgAMA1ΔDIIloop in complex with the EtRON2 peptide. When analyzed in the context of existing AMA1-RON2 structures, spatially distinct anchor points in the AMA1 groove were identified that, when engaged, appear to provide the necessary traction to outcompete the DII loop. Collectively, these data support a model where the AMA1 DII loop serves as a structural gatekeeper to selectively filter out ligands otherwise capable of binding with high affinity in the AMA1 apical groove. These data also highlight the importance of considering the functional implications of the DII loop in the ongoing development of therapeutic intervention strategies targeting the AMA1-RON2 invasion complex.
Organizational Affiliation:
Department of Biochemistry & Microbiology, University of Victoria, PO Box 3055 STN CSC, Victoria, BC, V8W 3P6, Canada.