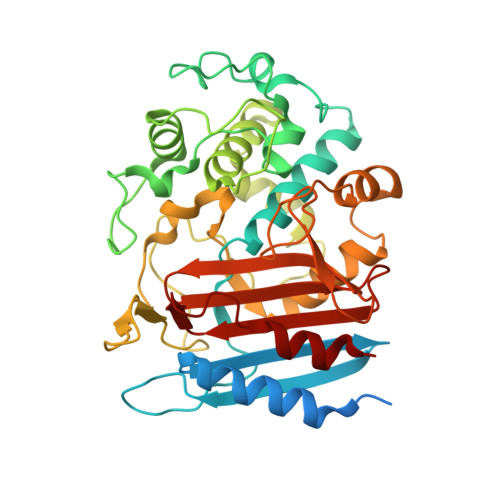
Structural and computational analysis of peptide recognition mechanism of class-C type penicillin binding protein, alkaline D-peptidase from Bacillus cereus DF4-B
Nakano, S., Okazaki, S., Ishitsubo, E., Kawahara, N., Komeda, H., Tokiwa, H., Asano, Y.(2015) Sci Rep 5: 13836-13836
- PubMed: 26370172
- DOI: https://doi.org/10.1038/srep13836
- Primary Citation of Related Structures:
4Y7P - PubMed Abstract:
Alkaline D-peptidase from Bacillus cereus DF4-B, called ADP, is a D-stereospecific endopeptidase reacting with oligopeptides containing D-phenylalanine (D-Phe) at N-terminal penultimate residue. ADP has attracted increasing attention because it is useful as a catalyst for synthesis of D-Phe oligopeptides or, with the help of substrate mimetics, L-amino acid peptides and proteins. Structure and functional analysis of ADP is expected to elucidate molecular mechanism of ADP. In this study, the crystal structure of ADP (apo) form was determined at 2.1 Å resolution. The fold of ADP is similar to that of the class C penicillin-binding proteins of type-AmpH. Docking simulations and fragment molecular orbital analyses of two peptides, (D-Phe)4 and (D-Phe)2-(L-Phe)2, with the putative substrate binding sites of ADP indicated that the P1 residue of the peptide interacts with hydrophobic residues at the S1 site of ADP. Furthermore, molecular dynamics simulation of ADP for 50 nsec suggested that the ADP forms large cavity at the active site. Formation of the cavity suggested that the ADP has open state in the solution. For the ADP, having the open state is convenient to bind the peptides having bulky side chain, such as (D-Phe)4. Taken together, we predicted peptide recognition mechanism of ADP.
Organizational Affiliation:
Biotechnology Research Center and Department of Biotechnology, Toyama Prefectural University, 5180 Kurokawa, Imizu, Toyama 939-0398, Japan.