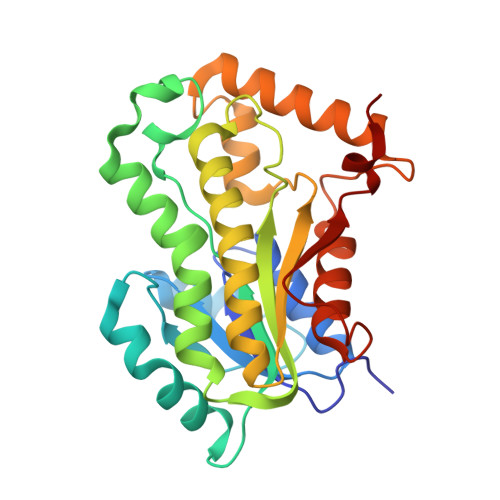
Crystal structure of the enoyl-ACP reductase of Mycobacterium tuberculosis (InhA) in the apo-form and in complex with the active metabolite of isoniazid pre-formed by a biomimetic approach.
Chollet, A., Mourey, L., Lherbet, C., Delbot, A., Julien, S., Baltas, M., Bernadou, J., Pratviel, G., Maveyraud, L., Bernardes-Genisson, V.(2015) J Struct Biol 190: 328-337
- PubMed: 25891098
- DOI: https://doi.org/10.1016/j.jsb.2015.04.008
- Primary Citation of Related Structures:
4TRM, 4TRN, 4TRO - PubMed Abstract:
InhA is an enoyl-ACP reductase of Mycobacterium tuberculosis implicated in the biosynthesis of mycolic acids, essential constituents of the mycobacterial cell wall. To date, this enzyme is considered as a promising target for the discovery of novel antitubercular drugs. In this work, we describe the first crystal structure of the apo form of the wild-type InhA at 1.80Å resolution as well as the crystal structure of InhA in complex with the synthetic metabolite of the antitubercular drug isoniazid refined to 1.40Å. This metabolite, synthesized in the absence of InhA, is able to displace and replace the cofactor NADH in the enzyme active site. This work provides a unique opportunity to enlighten the structural adaptation of apo-InhA to the binding of the NADH cofactor or of the isoniazid adduct. In addition, a differential scanning fluorimetry study of InhA, in the apo-form as well as in the presence of NAD(+), NADH and INH-NADH was performed showing that binding of the INH-NADH adduct had a strong stabilizing effect.
Organizational Affiliation:
Laboratoire de Chimie de Coordination (LCC), Centre National de la Recherche Scientifique (CNRS), 205 Route de Narbonne, BP 44099, F-31077 Toulouse, Cedex 4, France; Université de Toulouse, Université Paul Sabatier, INPT, F-31077 Toulouse, Cedex 4, France.