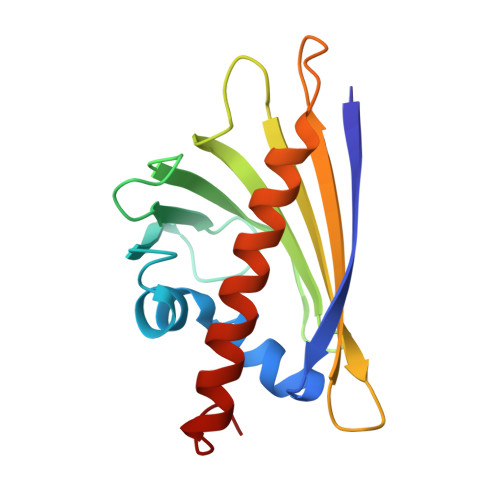
Ligand binding modulates the structural dynamics and compactness of the major birch pollen allergen
Grutsch, S., Fuchs, J.E., Freier, R., Kofler, S., Bibi, M., Asam, C., Wallner, M., Ferreira, F., Brandstetter, H., Liedl, K.R., Tollinger, M.(2014) Biophys J 107: 2963-2972
- PubMed: 25517162
- DOI: https://doi.org/10.1016/j.bpj.2014.10.062
- Primary Citation of Related Structures:
4QIP - PubMed Abstract:
Pathogenesis-related plant proteins of class-10 (PR-10) are essential for storage and transport of small molecules. A prominent member of the PR-10 family, the major birch pollen allergen Bet v 1, is the main cause of spring pollinosis in the temperate climate zone of the northern hemisphere. Bet v 1 binds various ligand molecules to its internal cavity, and immunologic effects of the presence of ligand have been discussed. However, the mechanism of binding has remained elusive. In this study, we show that in solution Bet v 1.0101 is conformationally heterogeneous and cannot be represented by a single structure. NMR relaxation data suggest that structural dynamics are fundamental for ligand access to the protein interior. Complex formation then leads to significant rigidification of the protein along with a compaction of its 3D structure. The data presented herein provide a structural basis for understanding the immunogenic and allergenic potential of ligand binding to Bet v 1 allergens.
Organizational Affiliation:
Institute of Organic Chemistry, University of Innsbruck, Innsbruck, Austria.