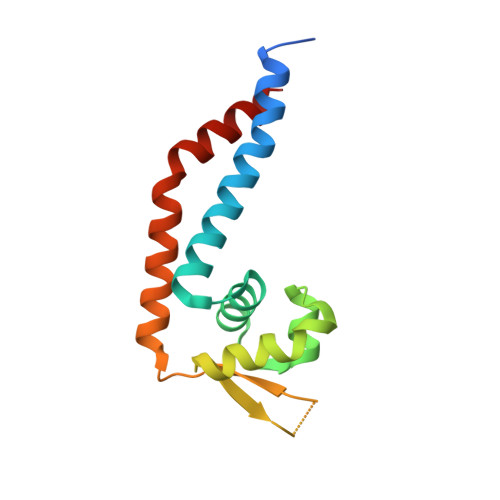
Structure of Rot, a global regulator of virulence genes in Staphylococcus aureus.
Zhu, Y., Fan, X., Zhang, X., Jiang, X., Niu, L., Teng, M., Li, X.(2014) Acta Crystallogr D Biol Crystallogr 70: 2467-2476
- PubMed: 25195759
- DOI: https://doi.org/10.1107/S1399004714015326
- Primary Citation of Related Structures:
4Q77 - PubMed Abstract:
Staphylococcus aureus is a highly versatile pathogen that can infect human tissue by producing a large arsenal of virulence factors that are tightly regulated by a complex regulatory network. Rot, which shares sequence similarity with SarA homologues, is a global regulator that regulates numerous virulence genes. However, the recognition model of Rot for the promoter region of target genes and the putative regulation mechanism remain elusive. In this study, the 1.77 Å resolution X-ray crystal structure of Rot is reported. The structure reveals that two Rot molecules form a compact homodimer, each of which contains a typical helix-turn-helix module and a β-hairpin motif connected by a flexible loop. Fluorescence polarization results indicate that Rot preferentially recognizes AT-rich dsDNA with ~30-base-pair nucleotides and that the conserved positively charged residues on the winged-helix motif are vital for binding to the AT-rich dsDNA. It is proposed that the DNA-recognition model of Rot may be similar to that of SarA, SarR and SarS, in which the helix-turn-helix motifs of each monomer interact with the major grooves of target dsDNA and the winged motifs contact the minor grooves. Interestingly, the structure shows that Rot adopts a novel dimerization model that differs from that of other SarA homologues. As expected, perturbation of the dimer interface abolishes the dsDNA-binding ability of Rot, suggesting that Rot functions as a dimer. In addition, the results have been further confirmed in vivo by measuring the transcriptional regulation of α-toxin, a major virulence factor produced by most S. aureus strains.
Organizational Affiliation:
Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, People's Republic of China.