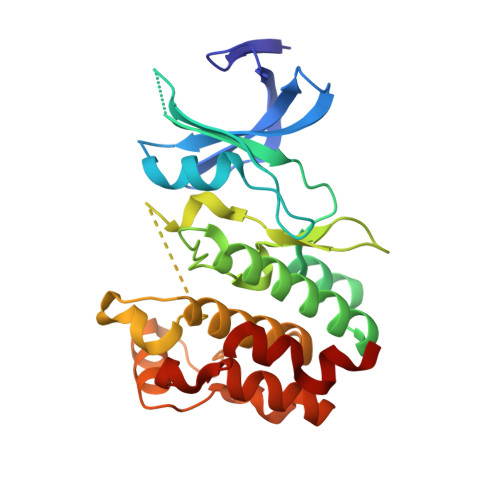
Chloride Sensing by WNK1 Involves Inhibition of Autophosphorylation.
Piala, A.T., Moon, T.M., Akella, R., He, H., Cobb, M.H., Goldsmith, E.J.(2014) Sci Signal 7: ra41-ra41
- PubMed: 24803536
- DOI: https://doi.org/10.1126/scisignal.2005050
- Primary Citation of Related Structures:
4PWN, 4Q2A - PubMed Abstract:
WNK1 [with no lysine (K)] is a serine-threonine kinase associated with a form of familial hypertension. WNK1 is at the top of a kinase cascade, leading to phosphorylation of several cotransporters, in particular those transporting sodium, potassium, and chloride (NKCC), sodium and chloride (NCC), and potassium and chloride (KCC). The responsiveness of NKCC, NCC, and KCC to changes in extracellular chloride parallels their phosphorylation state, provoking the proposal that these transporters are controlled by a chloride-sensitive protein kinase. We found that chloride stabilizes the inactive conformation of WNK1, preventing kinase autophosphorylation and activation. Crystallographic studies of inactive WNK1 in the presence of chloride revealed that chloride binds directly to the catalytic site, providing a basis for the unique position of the catalytic lysine. Mutagenesis of the chloride-binding site rendered the kinase less sensitive to inhibition of autophosphorylation by chloride, validating the binding site. Thus, these data suggest that WNK1 functions as a chloride sensor through direct binding of a regulatory chloride ion to the active site, which inhibits autophosphorylation.
Organizational Affiliation:
1Department of Biophysics, The University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, TX 75390, USA.