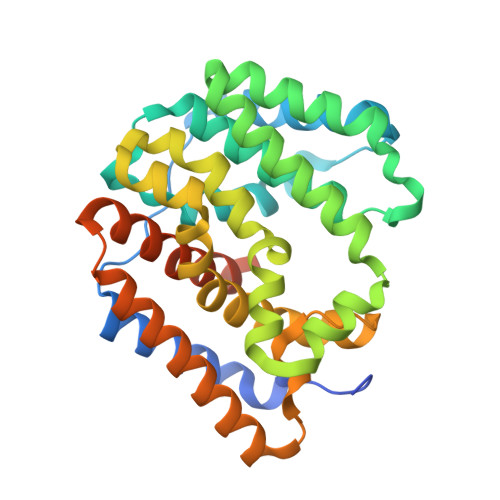
The first structure of a bacterial diterpene cyclase: CotB2.
Janke, R., Gorner, C., Hirte, M., Bruck, T., Loll, B.(2014) Acta Crystallogr D Biol Crystallogr 70: 1528-1537
- PubMed: 24914964
- DOI: https://doi.org/10.1107/S1399004714005513
- Primary Citation of Related Structures:
4OMG, 4OMH - PubMed Abstract:
Sesquiterpenes and diterpenes are a diverse class of secondary metabolites that are predominantly derived from plants and some prokaryotes. The properties of these natural products encompass antitumor, antibiotic and even insecticidal activities. Therefore, they are interesting commercial targets for the chemical and pharmaceutical industries. Owing to their structural complexity, these compounds are more efficiently accessed by metabolic engineering of microbial systems than by chemical synthesis. This work presents the first crystal structure of a bacterial diterpene cyclase, CotB2 from the soil bacterium Streptomyces melanosporofaciens, at 1.64 Å resolution. CotB2 is a diterpene cyclase that catalyzes the cyclization of the linear geranylgeranyl diphosphate to the tricyclic cyclooctat-9-en-7-ol. The subsequent oxidation of cyclooctat-9-en-7-ol by two cytochrome P450 monooxygenases leads to bioactive cyclooctatin. Plasticity residues that decorate the active site of CotB2 have been mutated, resulting in alternative monocyclic, dicyclic and tricyclic compounds that show bioactivity. These new compounds shed new light on diterpene cyclase reaction mechanisms. Furthermore, the product of mutant CotB2(W288G) produced the new antibiotic compound (1R,3E,7E,11S,12S)-3,7,18-dolabellatriene, which acts specifically against multidrug-resistant Staphylococcus aureus. This opens a sustainable route for the industrial-scale production of this bioactive compound.
Organizational Affiliation:
Institut für Chemie und Biochemie, Abteilung Strukturbiochemie, Freie Universität Berlin, Takustrasse 6, 14195 Berlin, Germany.