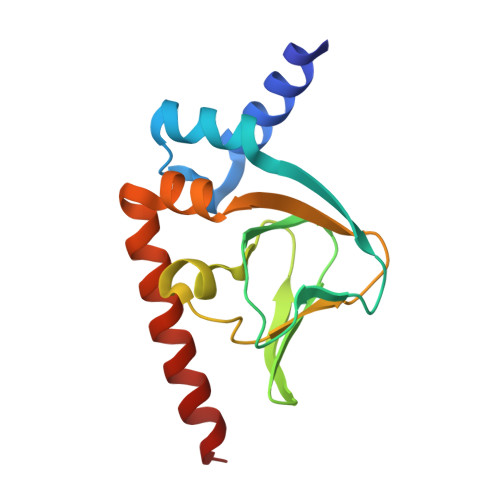
Crystal Structures of the Carboxyl cGMP Binding Domain of the Plasmodium falciparum cGMP-dependent Protein Kinase Reveal a Novel Capping Triad Crucial for Merozoite Egress.
Kim, J.J., Flueck, C., Franz, E., Sanabria-Figueroa, E., Thompson, E., Lorenz, R., Bertinetti, D., Baker, D.A., Herberg, F.W., Kim, C.(2015) PLoS Pathog 11: e1004639-e1004639
- PubMed: 25646845
- DOI: https://doi.org/10.1371/journal.ppat.1004639
- Primary Citation of Related Structures:
4OFF, 4OFG - PubMed Abstract:
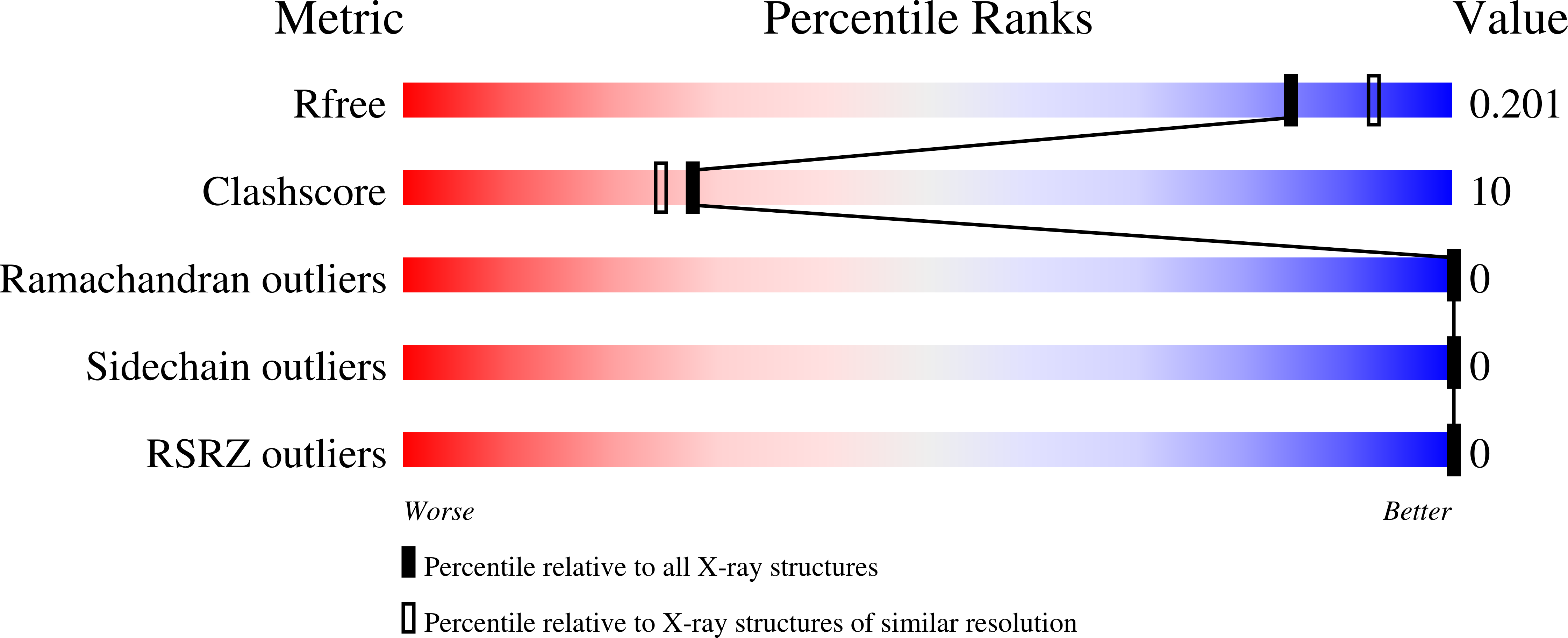
The Plasmodium falciparum cGMP-dependent protein kinase (PfPKG) is a key regulator across the malaria parasite life cycle. Little is known about PfPKG's activation mechanism. Here we report that the carboxyl cyclic nucleotide binding domain functions as a "gatekeeper" for activation by providing the highest cGMP affinity and selectivity. To understand the mechanism, we have solved its crystal structures with and without cGMP at 2.0 and 1.9 Å, respectively. These structures revealed a PfPKG-specific capping triad that forms upon cGMP binding, and disrupting the triad reduces kinase activity by 90%. Furthermore, mutating these residues in the parasite prevents blood stage merozoite egress, confirming the essential nature of the triad in the parasite. We propose a mechanism of activation where cGMP binding allosterically triggers the conformational change at the αC-helix, which bridges the regulatory and catalytic domains, causing the capping triad to form and stabilize the active conformation.
Organizational Affiliation:
Department of Pharmacology, Baylor College of Medicine, Houston, Texas, United States of America; Department of Biochemistry, University of Kassel, Kassel, Hesse, Germany.