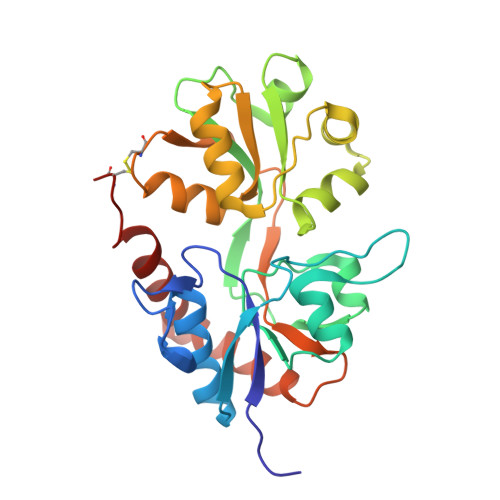
Synthesis, pharmacological and structural characterization, and thermodynamic aspects of GluA2-positive allosteric modulators with a 3,4-dihydro-2H-1,2,4-benzothiadiazine 1,1-dioxide scaffold.
Nrholm, A.B., Francotte, P., Olsen, L., Krintel, C., Frydenvang, K., Goffin, E., Challal, S., Danober, L., Botez-Pop, I., Lestage, P., Pirotte, B., Kastrup, J.S.(2013) J Med Chem 56: 8736-8745
- PubMed: 24131202
- DOI: https://doi.org/10.1021/jm4012092
- Primary Citation of Related Structures:
4N07 - PubMed Abstract:
Positive allosteric modulators of ionotropic glutamate receptors are potential compounds for treatment of cognitive disorders, e.g., Alzheimer's disease. The modulators bind within the dimer interface of the ligand-binding domain (LBD) and stabilize the agonist-bound conformation, thereby slowing receptor desensitization and/or deactivation. Here we describe the synthesis and pharmacological testing at GluA2 of a new generation of 3,4-dihydro-2H-1,2,4-benzothiadiazine 1,1-dioxides. The most potent modulator 3 in complex with GluA2-LBD-L483Y-N754S was subjected to structural analysis by X-ray crystallography, and the thermodynamics of binding was studied by isothermal titration calorimetry. Compound 3 binds to GluA2-LBD-L483Y-N754S with a Kd of 0.35 μM (ΔH = -7.5 kcal/mol and -TΔS = -1.3 kcal/mol). This is the first time that submicromolar binding affinity has been achieved for this type of positive allosteric modulator. The major structural factor increasing the binding affinity of 3 seems to be interactions between the cyclopropyl group of 3 and the backbone of Phe495 and Met496.
Organizational Affiliation:
Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen , Universitetsparken 2, DK-2100 Copenhagen, Denmark.