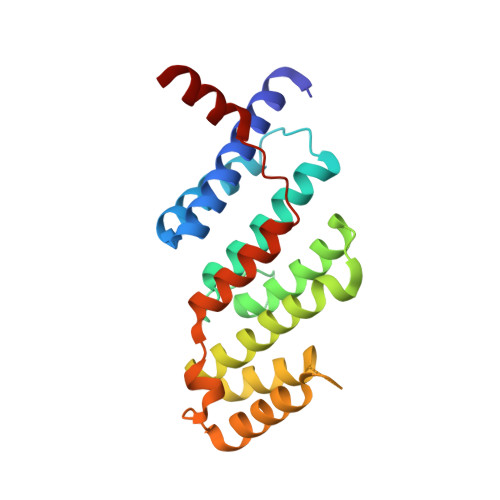
SLX4 Assembles a Telomere Maintenance Toolkit by Bridging Multiple Endonucleases with Telomeres
Wan, B., Yin, J., Horvath, K., Sarkar, J., Chen, Y., Wu, J., Wan, K., Lu, J., Gu, P., Yu, E.Y., Lue, N.F., Chang, S., Liu, Y., Lei, M.(2013) Cell Rep 4: 861-869
- PubMed: 24012755
- DOI: https://doi.org/10.1016/j.celrep.2013.08.017
- Primary Citation of Related Structures:
4M7C - PubMed Abstract:
SLX4 interacts with several endonucleases to resolve structural barriers in DNA metabolism. SLX4 also interacts with telomeric protein TRF2 in human cells. The molecular mechanism of these interactions at telomeres remains unknown. Here, we report the crystal structure of the TRF2-binding motif of SLX4 (SLX4TBM) in complex with the TRFH domain of TRF2 (TRF2TRFH) and map the interactions of SLX4 with endonucleases SLX1, XPF, and MUS81. TRF2 recognizes a unique HxLxP motif on SLX4 via the peptide-binding site in its TRFH domain. Telomeric localization of SLX4 and associated nucleases depend on the SLX4-endonuclease and SLX4-TRF2 interactions and the protein levels of SLX4 and TRF2. SLX4 assembles an endonuclease toolkit that negatively regulates telomere length via SLX1-catalyzed nucleolytic resolution of telomere DNA structures. We propose that the SLX4-TRF2 complex serves as a double-layer scaffold bridging multiple endonucleases with telomeres for recombination-based telomere maintenance.
Organizational Affiliation:
State Key Laboratory of Molecular Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yueyang Road, Shanghai 200031, China; National Center for Protein Science Shanghai, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yueyang Road, Shanghai 200031, China; Howard Hughes Medical Institute, University of Michigan Medical School, 1150 W. Medical Center Drive, Ann Arbor, MI 48109, USA; Department of Biological Chemistry, University of Michigan Medical School, 1150 W. Medical Center Drive, Ann Arbor, MI 48109, USA.