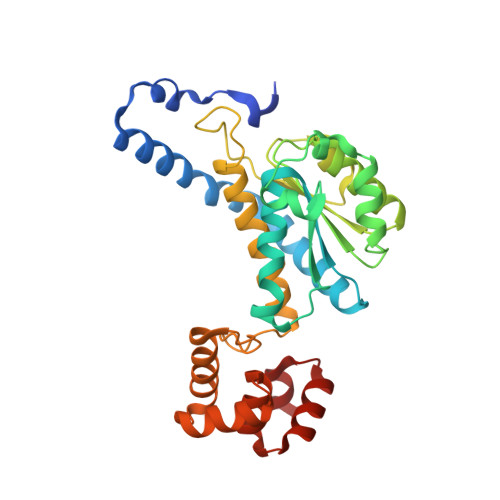
Structure of MurNAc 6-phosphate hydrolase (MurQ) from Haemophilus influenzae with a bound inhibitor.
Hadi, T., Hazra, S., Tanner, M.E., Blanchard, J.S.(2013) Biochemistry 52: 9358-9366
- PubMed: 24251551
- DOI: https://doi.org/10.1021/bi4010446
- Primary Citation of Related Structures:
4LZJ, 4M0D - PubMed Abstract:
The breakdown and recycling of peptidoglycan, an essential polymeric cell structure, occur in a number of bacterial species. A key enzyme in the recycling pathway of one of the components of the peptidoglycan layer, N-acetylmuramic acid (MurNAc), is MurNAc 6-phosphate hydrolase (MurQ). This enzyme catalyzes the cofactor-independent cleavage of a relatively nonlabile ether bond and presents an interesting target for mechanistic studies. Open chain product and substrate analogues were synthesized and tested as competitive inhibitors (K(is) values of 1.1 ± 0.3 and 0.23 ± 0.02 mM, respectively) of the MurNAc 6P hydrolase from Escherichia coli (MurQ-EC). To identify the roles of active site residues that are important for catalysis, the substrate analogue was cocrystallized with the MurNAc 6P hydrolase from Haemophilus influenzae (MurQ-HI) that was amenable to crystallographic studies. The cocrystal structure of MurQ-HI with the substrate analogue showed that Glu89 was located in the proximity of both the C2 atom and the oxygen at the C3 position of the bound inhibitor and that no other potential acid/base residue that could act as an active site acid/base was located in the vicinity. The conserved residues Glu120 and Lys239 were found within hydrogen bonding distance of the C5 hydroxyl group and C6 phosphate group, suggesting that they play a role in substrate binding and ring opening. Combining these results with previous biochemical data, we propose a one-base mechanism of action in which Glu89 functions to both deprotonate at the C2 position and assist in the departure of the lactyl ether at the C3 position. This same residue would serve to deprotonate the incoming water and reprotonate the enolate in the second half of the catalytic cycle.
Organizational Affiliation:
Department of Biochemistry, Albert Einstein College of Medicine , 1300 Morris Park Avenue, Bronx, New York 10461, United States.