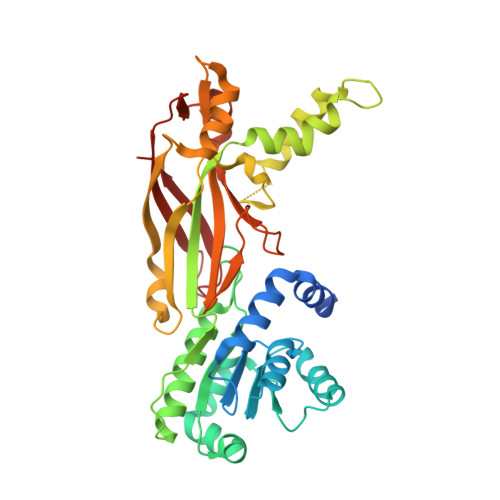
Crystal Structure of Arginine Methyltransferase 6 from Trypanosoma brucei
Wang, C., Zhu, Y., Chen, J., Li, X., Peng, J., Chen, J., Zou, Y., Zhang, Z., Jin, H., Yang, P., Wu, J., Niu, L., Gong, Q., Teng, M., Shi, Y.(2014) PLoS One 9: e87267-e87267
- PubMed: 24498306
- DOI: https://doi.org/10.1371/journal.pone.0087267
- Primary Citation of Related Structures:
4LWO, 4LWP - PubMed Abstract:
Arginine methylation plays vital roles in the cellular functions of the protozoan Trypanosoma brucei. The T. brucei arginine methyltransferase 6 (TbPRMT6) is a type I arginine methyltransferase homologous to human PRMT6. In this study, we report the crystal structures of apo-TbPRMT6 and its complex with the reaction product S-adenosyl-homocysteine (SAH). The structure of apo-TbPRMT6 displays several features that are different from those of type I PRMTs that were structurally characterized previously, including four stretches of insertion, the absence of strand β15, and a distinct dimerization arm. The comparison of the apo-TbPRMT6 and SAH-TbPRMT6 structures revealed the fine rearrangements in the active site upon SAH binding. The isothermal titration calorimetry results demonstrated that SAH binding greatly increases the affinity of TbPRMT6 to a substrate peptide derived from bovine histone H4. The western blotting and mass spectrometry results revealed that TbPRMT6 methylates bovine histone H4 tail at arginine 3 but cannot methylate several T. brucei histone tails. In summary, our results highlight the structural differences between TbPRMT6 and other type I PRMTs and reveal that the active site rearrangement upon SAH binding is important for the substrate binding of TbPRMT6.
Organizational Affiliation:
Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Anhui, China.