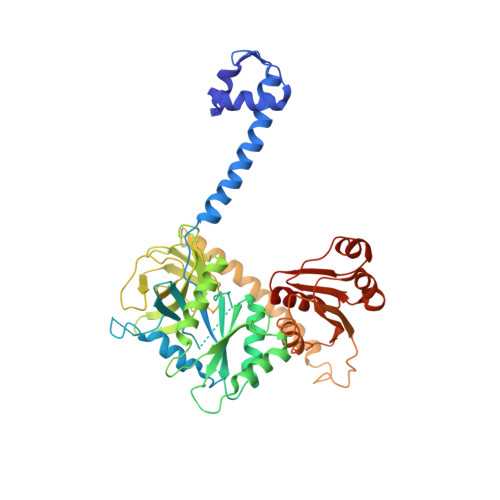
Initiation factor 2 crystal structure reveals a different domain organization from eukaryotic initiation factor 5B and mechanism among translational GTPases.
Eiler, D., Lin, J., Simonetti, A., Klaholz, B.P., Steitz, T.A.(2013) Proc Natl Acad Sci U S A 110: 15662-15667
- PubMed: 24029018
- DOI: https://doi.org/10.1073/pnas.1309360110
- Primary Citation of Related Structures:
4KJZ - PubMed Abstract:
The initiation of protein synthesis uses initiation factor 2 (IF2) in prokaryotes and a related protein named eukaryotic initiation factor 5B (eIF5B) in eukaryotes. IF2 is a GTPase that positions the initiator tRNA on the 30S ribosomal initiation complex and stimulates its assembly to the 50S ribosomal subunit to make the 70S ribosome. The 3.1-Å resolution X-ray crystal structures of the full-length Thermus thermophilus apo IF2 and its complex with GDP presented here exhibit two different conformations (all of its domains except C2 domain are visible). Unlike all other translational GTPases, IF2 does not have an effecter domain that stably contacts the switch II region of the GTPase domain. The domain organization of IF2 is inconsistent with the "articulated lever" mechanism of communication between the GTPase and initiator tRNA binding domains that has been proposed for eIF5B. Previous cryo-electron microscopy reconstructions, NMR experiments, and this structure show that IF2 transitions from being flexible in solution to an extended conformation when interacting with ribosomal complexes.
Organizational Affiliation:
Departments of Chemistry and Molecular Biochemistry and Biophysics, Yale University, New Haven, CT 06520.