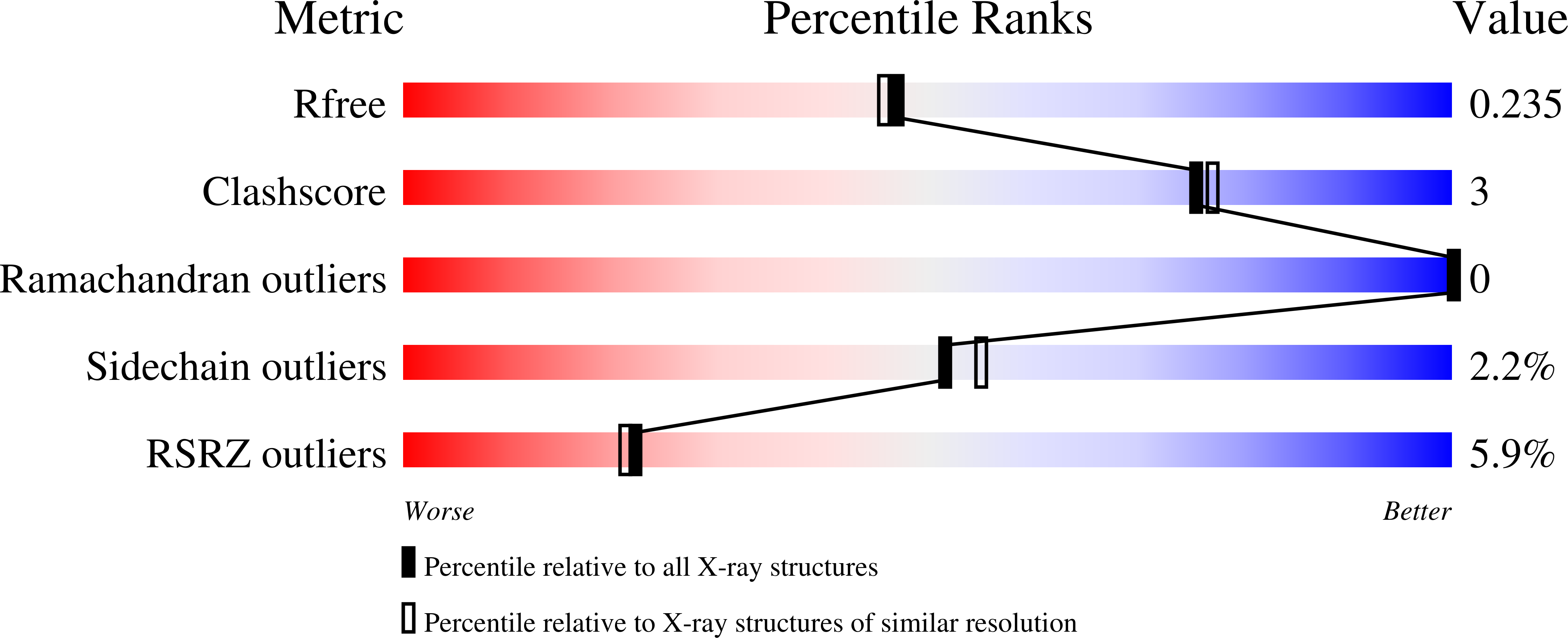
Crystal structure of the Staphylococcus aureus protein (NP_646141.1, domain 3912-4037) similar to streptococcal adhesins emb and ebhA/ebhB
Cymborowski, M., Shabalin, I.G., Joachimiak, G., Chruszcz, M., Gornicki, P., Zhang, R., Joachimiak, A., Minor, W.To be published.