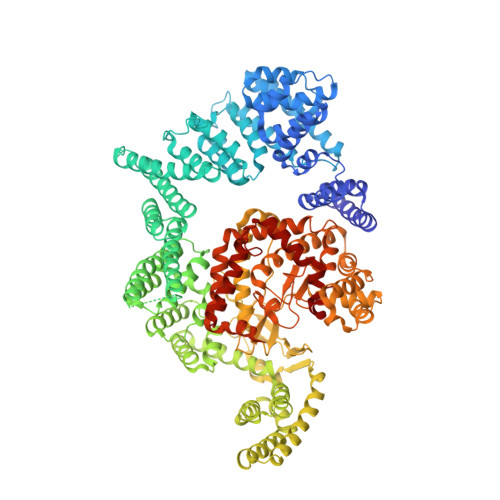
mTOR kinase structure, mechanism and regulation.
Yang, H., Rudge, D.G., Koos, J.D., Vaidialingam, B., Yang, H.J., Pavletich, N.P.(2013) Nature 497: 217-223
- PubMed: 23636326
- DOI: https://doi.org/10.1038/nature12122
- Primary Citation of Related Structures:
4JSN, 4JSP, 4JSV, 4JSX, 4JT5, 4JT6 - PubMed Abstract:
The mammalian target of rapamycin (mTOR), a phosphoinositide 3-kinase-related protein kinase, controls cell growth in response to nutrients and growth factors and is frequently deregulated in cancer. Here we report co-crystal structures of a complex of truncated mTOR and mammalian lethal with SEC13 protein 8 (mLST8) with an ATP transition state mimic and with ATP-site inhibitors. The structures reveal an intrinsically active kinase conformation, with catalytic residues and a catalytic mechanism remarkably similar to canonical protein kinases. The active site is highly recessed owing to the FKBP12-rapamycin-binding (FRB) domain and an inhibitory helix protruding from the catalytic cleft. mTOR-activating mutations map to the structural framework that holds these elements in place, indicating that the kinase is controlled by restricted access. In vitro biochemistry shows that the FRB domain acts as a gatekeeper, with its rapamycin-binding site interacting with substrates to grant them access to the restricted active site. Rapamycin-FKBP12 inhibits the kinase by directly blocking substrate recruitment and by further restricting active-site access. The structures also reveal active-site residues and conformational changes that underlie inhibitor potency and specificity.
Organizational Affiliation:
Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York 10065, USA.