2.06 Angstrom resolution crystal structure of phosphomethylpyrimidine kinase (thiD)from Clostridium difficile 630
Halavaty, A.S., Wawrzak, Z., Onopriyenko, O., Grimshaw, S., Savchenko, A., Anderson, W.F.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Phosphomethylpyrimidine kinase | 267 | Clostridioides difficile 630 | Mutation(s): 0 Gene Names: CD630_15990, thiD EC: 2.7.4.7 | 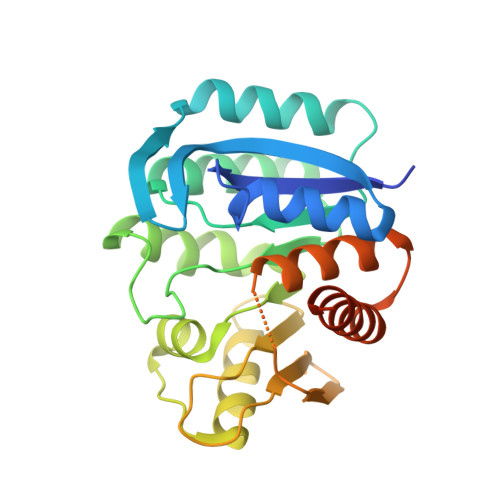 | |
UniProt | |||||
Find proteins for Q186F5 (Clostridioides difficile (strain 630)) Explore Q186F5 Go to UniProtKB: Q186F5 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q186F5 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| EPE Query on EPE | C [auth B] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 56.751 | α = 90 |
| b = 79.741 | β = 90 |
| c = 92.863 | γ = 90 |
| Software Name | Purpose |
|---|---|
| Blu-Ice | data collection |
| BALBES | phasing |
| REFMAC | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |