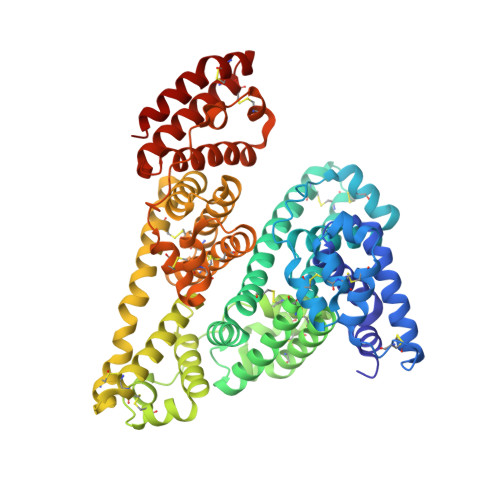
Crystallographic studies of the complexes of bovine and equine serum albumin with 3,5-diiodosalicylic acid.
Sekula, B., Zielinski, K., Bujacz, A.(2013) Int J Biol Macromol 60C: 316-324
- PubMed: 23769932
- DOI: https://doi.org/10.1016/j.ijbiomac.2013.06.004
- Primary Citation of Related Structures:
4J2V, 4JK4 - PubMed Abstract:
Due to their extraordinary binding properties, serum albumins are the main transporters of many small molecules in the circulatory system. Although all mammalian serum albumins exhibit quite high sequence similarity, their binding abilities are not the same. Until now, only human serum albumin (HSA) was subjected to extensive structural studies in complexes with various ligands. Here we present two crystal structures of the complexes of equine and bovine serum albumins with 3,5-diiodosalicylic acid (DIS), at resolutions 2.12 Å and 2.65 Å, respectively, and analyze interactions of the DIS ligand with both macromolecules. We highlight the differences in distribution of DIS binding sites between the bovine and equine serum albumins and compare results with the HSA binding ability of DIS and other structurally similar ligands.
Organizational Affiliation:
Institute of Technical Biochemistry, Lodz University of Technology, Stefanowskiego 4/10, 90-924 Lodz, Poland.