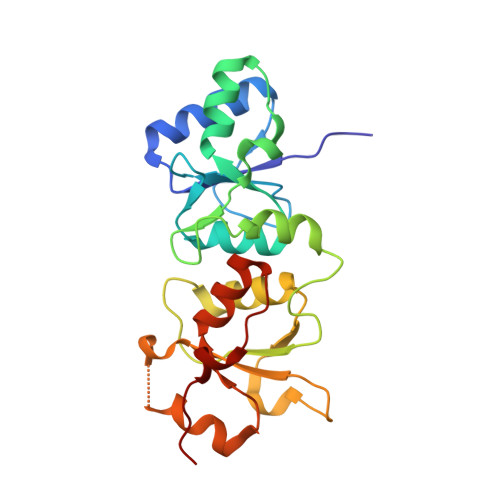
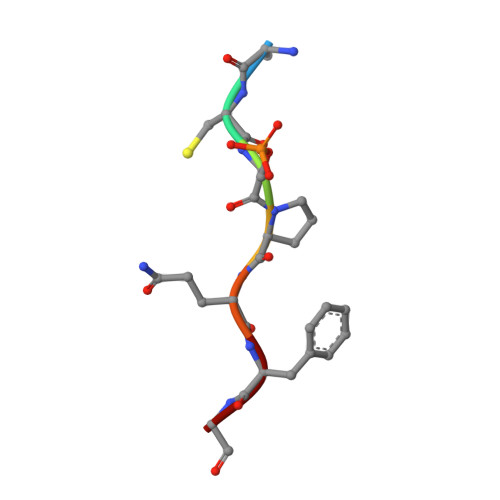
Structural Basis for the BRCA1 BRCT Interaction with the Proteins ATRIP and BAAT1.
Liu, X., Ladias, J.A.(2013) Biochemistry 52: 7618-7627
- PubMed: 24073851
- DOI: https://doi.org/10.1021/bi400714v
- Primary Citation of Related Structures:
4IFI, 4IGK - PubMed Abstract:
The breast and ovarian cancer susceptibility protein 1 (BRCA1) plays a central role in DNA damage response (DDR). Two tandem BRCA1 C-terminal (BRCT) domains interact with several proteins that function in DDR and contain the generally accepted motif pS-X-X-F (pS denoting phosphoserine and X any amino acid), including the ATR-interacting protein (ATRIP) and the BRCA1-associated protein required for ATM activation-1 (BAAT1). The crystal structures of the BRCA1 BRCTs bound to the phosphopeptides ATRIP (235-PEACpSPQFG-243) and BAAT1 (266-VARpSPVFSS-274) were determined at 1.75 Å and 2.2 Å resolution, respectively. The pSer and Phe(+3) anchor the phosphopeptides into the BRCT binding groove, with adjacent peptide residues contributing to the interaction. In the BRCA1-ATRIP structure, Gln(+2) is accommodated through a conformational change of the BRCA1 E1698 side chain. Importantly, isothermal titration calorimetry experiments showed that the size and charge of the side chains at peptide positions +1 and +2 contribute significantly to the BRCA1 BRCT-peptide binding affinity. In particular, the Asp(+1) and Glu(+2) in the human CDC27 peptide 816-HAAEpSDEF-823 abrogate the interaction with the BRCA1 BRCTs due in large part to electrostatic repulsion between Glu(+2) and E1698, indicating a preference of these domains for specific side chains at positions +1 and +2. These results emphasize the need for a systematic assessment of the contribution of the peptide residues surrounding pSer and Phe(+3) to the binding affinity and specificity of the BRCA1 BRCTs in order to elucidate the molecular mechanisms underlying the hierarchy of target selection by these versatile domains during DDR and tumorigenesis.
Organizational Affiliation:
Molecular Medicine Laboratory and Macromolecular Crystallography Unit, Department of Medicine, Harvard Medical School , Boston Massachusetts 02215, United States.