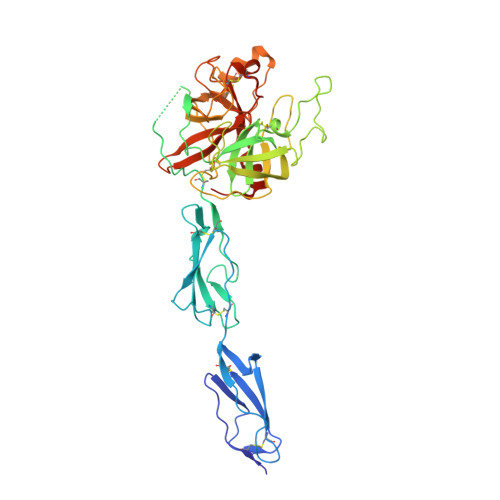
Quantitative characterization of the activation steps of mannan-binding lectin (MBL)-associated serine proteases (MASPs) points to the central role of MASP-1 in the initiation of the complement lectin pathway
Megyeri, M., Harmat, V., Major, B., Vegh, A., Balczer, J., Heja, D., Szilagyi, K., Datz, D., Pal, G., Zavodszky, P., Gal, P., Dobo, J.(2013) J Biol Chem 288: 8922-8934
- PubMed: 23386610
- DOI: https://doi.org/10.1074/jbc.M112.446500
- Primary Citation of Related Structures:
4IGD - PubMed Abstract:
Mannan-binding lectin (MBL)-associated serine proteases, MASP-1 and MASP-2, have been thought to autoactivate when MBL/ficolin·MASP complexes bind to pathogens triggering the complement lectin pathway. Autoactivation of MASPs occurs in two steps: 1) zymogen autoactivation, when one proenzyme cleaves another proenzyme molecule of the same protease, and 2) autocatalytic activation, when the activated protease cleaves its own zymogen. Using recombinant catalytic fragments, we demonstrated that a stable proenzyme MASP-1 variant (R448Q) cleaved the inactive, catalytic site Ser-to-Ala variant (S646A). The autoactivation steps of MASP-1 were separately quantified using these mutants and the wild type enzyme. Analogous mutants were made for MASP-2, and rate constants of the autoactivation steps as well as the possible cross-activation steps between MASP-1 and MASP-2 were determined. Based on the rate constants, a kinetic model of lectin pathway activation was outlined. The zymogen autoactivation rate of MASP-1 is ∼3000-fold higher, and the autocatalytic activation of MASP-1 is about 140-fold faster than those of MASP-2. Moreover, both activated and proenzyme MASP-1 can effectively cleave proenzyme MASP-2. MASP-3, which does not autoactivate, is also cleaved by MASP-1 quite efficiently. The structure of the catalytic region of proenzyme MASP-1 R448Q was solved at 2.5 Å. Proenzyme MASP-1 R448Q readily cleaves synthetic substrates, and it is inhibited by a specific canonical inhibitor developed against active MASP-1, indicating that zymogen MASP-1 fluctuates between an inactive and an active-like conformation. The determined structure provides a feasible explanation for this phenomenon. In summary, autoactivation of MASP-1 is crucial for the activation of MBL/ficolin·MASP complexes, and in the proenzymic phase zymogen MASP-1 controls the process.
Organizational Affiliation:
Institute of Enzymology, Research Centre for Natural Sciences, Hungarian Academy of Sciences, 29 Karolina Street, H-1113 Budapest, Hungary.