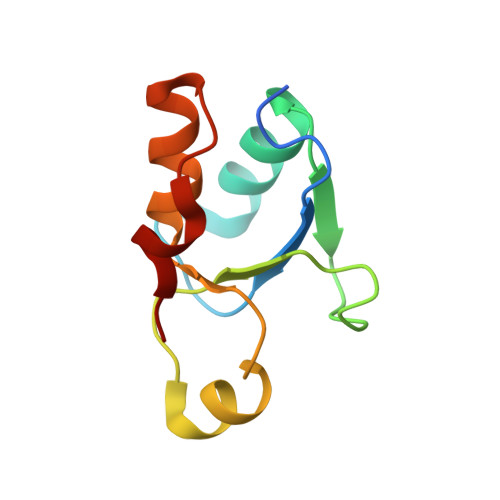
Structure and Functional Analysis of the BRCT Domain of Translesion Synthesis DNA Polymerase Rev1.
Pryor, J.M., Gakhar, L., Washington, M.T.(2013) Biochemistry 52: 254-263
- PubMed: 23240687
- DOI: https://doi.org/10.1021/bi301572z
- Primary Citation of Related Structures:
4ID3 - PubMed Abstract:
Translesion synthesis (TLS) is a pathway in which specialized, low-fidelity DNA polymerases are used to overcome replication blocks caused by DNA damage. The use of this pathway often results in somatic mutations that can drive carcinogenesis. Rev1 is a TLS polymerase found in all eukaryotes that plays a pivotal role in mediating DNA damage-induced mutagenesis. It possesses a BRCA1 C-terminal (BRCT) domain that is required for its function. The rev1-1 allele encodes a mutant form of Rev1 with a G193R substitution in this domain, which reduces the level of DNA damage-induced mutagenesis. Despite its clear importance in mutagenic TLS, the role of the BRCT domain is unknown. Here, we report the X-ray crystal structure of the yeast Rev1 BRCT domain and show that substitutions in residues constituting its phosphate-binding pocket do not affect mutagenic TLS. This suggests that the role of the Rev1 BRCT domain is not to recognize phosphate groups on protein binding partners or on DNA. We also found that residue G193 is located in a conserved turn region of the BRCT domain, and our in vivo and in vitro studies suggest that the G193R substitution may disrupt Rev1 function by destabilizing the fold of the BRCT domain.
Organizational Affiliation:
Department of Biochemistry, Carver College of Medicine, University of Iowa, Iowa City, IA 52242, USA.