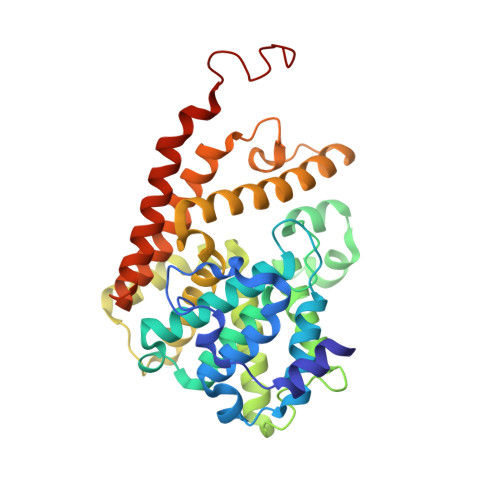
X-ray Crystal Structure of Phosphodiesterase 2 in Complex with a Highly Selective, Nanomolar Inhibitor Reveals a Binding-Induced Pocket Important for Selectivity.
Zhu, J., Yang, Q., Dai, D., Huang, Q.(2013) J Am Chem Soc 135: 11708-11711
- PubMed: 23899287
- DOI: https://doi.org/10.1021/ja404449g
- Primary Citation of Related Structures:
4HTX, 4HTZ - PubMed Abstract:
To better understand the structural origins of inhibitor selectivity of human phosphodieasterase families (PDEs 1-11), here we report the X-ray crystal structure of PDE2 in complex with a highly selective, nanomolar inhibitor (BAY60-7550) at 1.9 Å resolution, and the structure of apo PDE2 at 2.0 Å resolution. The crystal structures reveal that the inhibitor binds to the PDE2 active site by using not only the conserved glutamine-switch mechanism for substrate binding, but also a binding-induced, hydrophobic pocket that was not reported previously. In silico affinity profiling by molecular docking indicates that the inhibitor binding to this pocket contributes significantly to the binding affinity and thereby improves the inhibitor selectivity for PDE2. Our results highlight a structure-based design strategy that exploits the potential binding-induced pockets to achieve higher selectivity in the PDE inhibitor development.
Organizational Affiliation:
State Key Laboratory of Genetic Engineering, School of Life Sciences, Fudan University, Shanghai 200433, China.