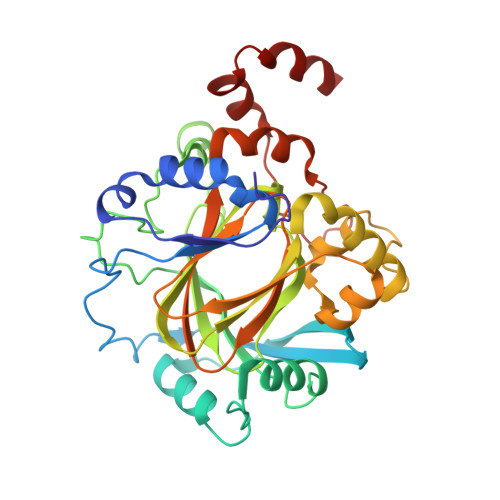
Structural and Functional Analysis of JMJD2D Reveals Molecular Basis for Site-Specific Demethylation among JMJD2 Demethylases.
Krishnan, S., Trievel, R.C.(2013) Structure 21: 98-108
- PubMed: 23219879
- DOI: https://doi.org/10.1016/j.str.2012.10.018
- Primary Citation of Related Structures:
4HON, 4HOO - PubMed Abstract:
JMJD2 lysine demethylases (KDMs) participate in diverse genomic processes. Most JMJD2 homologs display dual selectivity toward H3K9me3 and H3K36me3, with the exception of JMJD2D, which is specific for H3K9me3. Here, we report the crystal structures of the JMJD2D⋅2-oxoglutarate⋅H3K9me3 ternary complex and JMJD2D apoenzyme. Utilizing structural alignments with JMJD2A, molecular docking, and kinetic analysis with an array of histone peptide substrates, we elucidate the specific signatures that permit efficient recognition of H3K9me3 by JMJD2A and JMJD2D, and the residues in JMJD2D that occlude H3K36me3 demethylation. Surprisingly, these results reveal that JMJD2A and JMJD2D exhibit subtle yet important differences in H3K9me3 recognition, despite the overall similarity in the substrate-binding conformation. Further, we show that H3T11 phosphorylation abrogates demethylation by JMJD2 KDMs. Together, these studies reveal the molecular basis for JMJD2 site specificity and provide a framework for structure-based design of selective inhibitors of JMJD2 KDMs implicated in disease.
Organizational Affiliation:
Department of Biological Chemistry, 1150 West Medical Center Drive, 5301 Medical Science Research Building III, University of Michigan, Ann Arbor, MI 48109, USA.