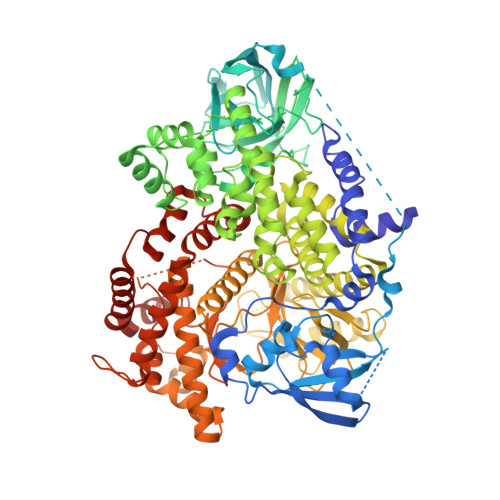
Cis-Amide isosteric replacement in thienobenzoxepin inhibitors of PI3-kinase.
Staben, S.T., Blaquiere, N., Tsui, V., Kolesnikov, A., Do, S., Bradley, E.K., Dotson, J., Goldsmith, R., Heffron, T.P., Lesnick, J., Lewis, C., Murray, J., Nonomiya, J., Olivero, A.G., Pang, J., Rouge, L., Salphati, L., Wei, B., Wiesmann, C., Wu, P.(2013) Bioorg Med Chem Lett 23: 897-901
- PubMed: 23265894
- DOI: https://doi.org/10.1016/j.bmcl.2012.10.121
- Primary Citation of Related Structures:
4HLE - PubMed Abstract:
Substructural class effects surrounding replacement of a 'cis' N-methyl aniline amide within potent and selective thienobenzoxepin PI3-kinase inhibitors are disclosed. While a simple aryl to alkyl switch was not tolerated due to differences in preferred amide conformation, heterocyclic amide isosteres with maintained aryl substitution improved potency and metabolic stability at the cost of physical properties. These gains in potency allowed lipophilic deconstruction of the arene to simple branched alkyl substituents. As such, overall lipophilicity-neutral, MW decreases were realized relative to the aniline amide series. The improved properties for lead compound 21 resulted in high permeability, solubility and bioavailability.
Organizational Affiliation:
Discovery Chemistry, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA. stevents@gene.com