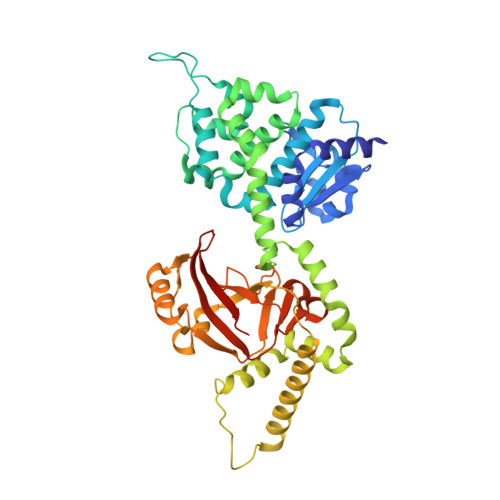
Certhrax Toxin, an Anthrax-related ADP-ribosyltransferase from Bacillus cereus.
Visschedyk, D., Rochon, A., Tempel, W., Dimov, S., Park, H.W., Merrill, A.R.(2012) J Biol Chem 287: 41089-41102
- PubMed: 22992735
- DOI: https://doi.org/10.1074/jbc.M112.412809
- Primary Citation of Related Structures:
4FK7, 4FXQ, 4GF1 - PubMed Abstract:
We identified Certhrax, the first anthrax-like mART toxin from the pathogenic G9241 strain of Bacillus cereus. Certhrax shares 31% sequence identity with anthrax lethal factor from Bacillus anthracis; however, we have shown that the toxicity of Certhrax resides in the mART domain, whereas anthrax uses a metalloprotease mechanism. Like anthrax lethal factor, Certhrax was found to require protective antigen for host cell entry. This two-domain enzyme was shown to be 60-fold more toxic to mammalian cells than anthrax lethal factor. Certhrax localizes to distinct regions within mouse RAW264.7 cells by 10 min postinfection and is extranuclear in its cellular location. Substitution of catalytic residues shows that the mART function is responsible for the toxicity, and it binds NAD(+) with high affinity (K(D) = 52.3 ± 12.2 μM). We report the 2.2 Å Certhrax structure, highlighting its structural similarities and differences with anthrax lethal factor. We also determined the crystal structures of two good inhibitors (P6 (K(D) = 1.7 ± 0.2 μM, K(i) = 1.8 ± 0.4 μM) and PJ34 (K(D) = 5.8 ± 2.6 μM, K(i) = 9.6 ± 0.3 μM)) in complex with Certhrax. As with other toxins in this family, the phosphate-nicotinamide loop moves toward the NAD(+) binding site with bound inhibitor. These results indicate that Certhrax may be important in the pathogenesis of B. cereus.
Organizational Affiliation:
Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario N1G 2W1, Canada.