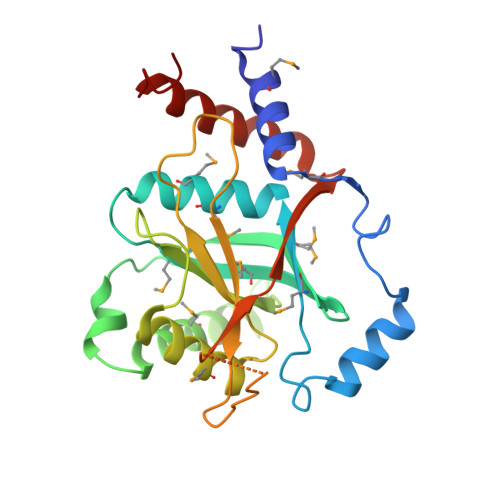
Insights into the regulation of the human COP9 signalosome catalytic subunit, CSN5/Jab1.
Echalier, A., Pan, Y., Birol, M., Tavernier, N., Pintard, L., Hoh, F., Ebel, C., Galophe, N., Claret, F.X., Dumas, C.(2013) Proc Natl Acad Sci U S A 110: 1273-1278
- PubMed: 23288897
- DOI: https://doi.org/10.1073/pnas.1209345110
- Primary Citation of Related Structures:
4F7O - PubMed Abstract:
The COP9 (Constitutive photomorphogenesis 9) signalosome (CSN), a large multiprotein complex that resembles the 19S lid of the 26S proteasome, plays a central role in the regulation of the E3-cullin RING ubiquitin ligases (CRLs). The catalytic activity of the CSN complex, carried by subunit 5 (CSN5/Jab1), resides in the deneddylation of the CRLs that is the hydrolysis of the cullin-neural precursor cell expressed developmentally downregulated gene 8 (Nedd8)isopeptide bond. Whereas CSN-dependent CSN5 displays isopeptidase activity, it is intrinsically inactive in other physiologically relevant forms. Here we analyze the crystal structure of CSN5 in its catalytically inactive form to illuminate the molecular basis for its activation state. We show that CSN5 presents a catalytic domain that brings essential elements to understand its activity control. Although the CSN5 active site is catalytically competent and compatible with di-isopeptide binding, the Ins-1 segment obstructs access to its substrate-binding site, and structural rearrangements are necessary for the Nedd8-binding pocket formation. Detailed study of CSN5 by molecular dynamics unveils signs of flexibility and plasticity of the Ins-1 segment. These analyses led to the identification of a molecular trigger implicated in the active/inactive switch that is sufficient to impose on CSN5 an active isopeptidase state. We show that a single mutation in the Ins-1 segment restores biologically relevant deneddylase activity. This study presents detailed insights into CSN5 regulation. Additionally, a dynamic monomer-dimer equilibrium exists both in vitro and in vivo and may be functionally relevant.
Organizational Affiliation:
Centre de Biochimie Structurale, Centre National de la Recherche Scientifique, Institut National de la Santé et de la Recherche Médicale-Unités Mixtes de Recherche 5048 and 1054, Université Montpellier I, 34090 Montpellier, France.