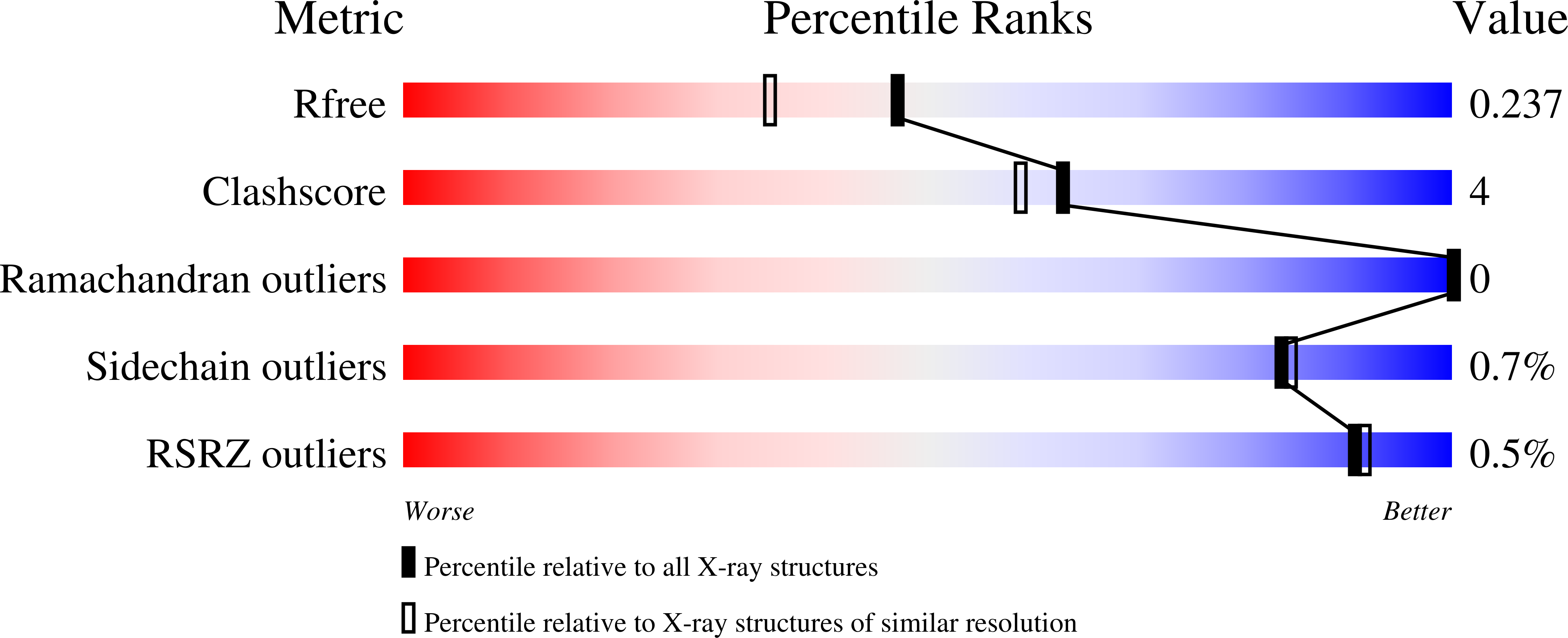
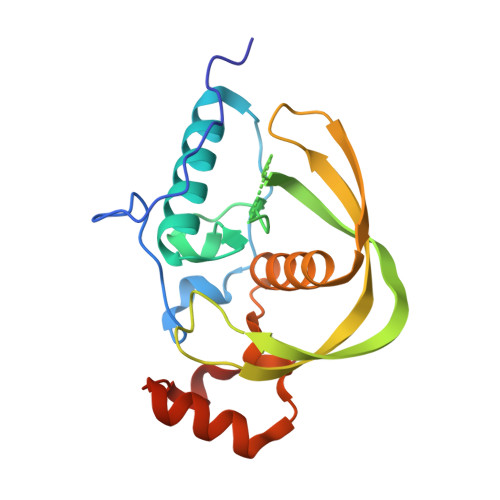
Structure and function of a cyanophage-encoded peptide deformylase.
Frank, J.A., Lorimer, D., Youle, M., Witte, P., Craig, T., Abendroth, J., Rohwer, F., Edwards, R.A., Segall, A.M., Burgin, A.B.(2013) ISME J 7: 1150-1160
- PubMed: 23407310
- DOI: https://doi.org/10.1038/ismej.2013.4
- Primary Citation of Related Structures:
3UWA, 3UWB, 4DR8, 4DR9 - PubMed Abstract:
Bacteriophages encode auxiliary metabolic genes that support more efficient phage replication. For example, cyanophages carry several genes to maintain host photosynthesis throughout infection, shuttling the energy and reducing power generated away from carbon fixation and into anabolic pathways. Photodamage to the D1/D2 proteins at the core of photosystem II necessitates their continual replacement. Synthesis of functional proteins in bacteria requires co-translational removal of the N-terminal formyl group by a peptide deformylase (PDF). Analysis of marine metagenomes to identify phage-encoded homologs of known metabolic genes found that marine phages carry PDF genes, suggesting that their expression during infection might benefit phage replication. We identified a PDF homolog in the genome of Synechococcus cyanophage S-SSM7. Sequence analysis confirmed that it possesses the three absolutely conserved motifs that form the active site in PDF metalloproteases. Phylogenetic analysis placed it within the Type 1B subclass, most closely related to the Arabidopsis chloroplast PDF, but lacking the C-terminal α-helix characteristic of that group. PDF proteins from this phage and from Synechococcus elongatus were expressed and characterized. The phage PDF is the more active enzyme and deformylates the N-terminal tetrapeptides from D1 proteins more efficiently than those from ribosomal proteins. Solution of the X-ray/crystal structures of those two PDFs to 1.95 Å resolution revealed active sites identical to that of the Type 1B Arabidopsis chloroplast PDF. Taken together, these findings show that many cyanophages encode a PDF with a D1 substrate preference that adds to the repertoire of genes used by phages to maintain photosynthetic activities.
Organizational Affiliation:
Department of Biology, San Diego State University, San Diego, CA 92182-7720, USA. jeralfrank@gmail.com