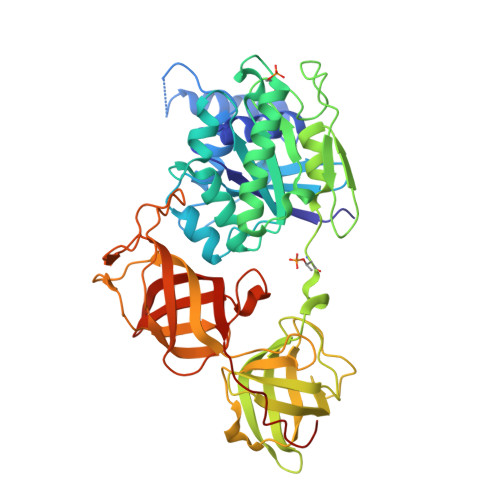
Mammalian Translation Elongation Factor Eef1A2: X-Ray Structure and New Features of Gdp/GTP Exchange Mechanism in Higher Eukaryotes
Crepin, T., Mccarthy, A., Negrutskii, A., Shalak, V.F., Tukalo, V., Vlasenko, D.O., Yaremchuk, A.D.(2014) Nucleic Acids Res 42: 12939
- PubMed: 25326326
- DOI: https://doi.org/10.1093/nar/gku974
- Primary Citation of Related Structures:
4C0S - PubMed Abstract:
Eukaryotic elongation factor eEF1A transits between the GTP- and GDP-bound conformations during the ribosomal polypeptide chain elongation. eEF1A*GTP establishes a complex with the aminoacyl-tRNA in the A site of the 80S ribosome. Correct codon-anticodon recognition triggers GTP hydrolysis, with subsequent dissociation of eEF1A*GDP from the ribosome. The structures of both the 'GTP'- and 'GDP'-bound conformations of eEF1A are unknown. Thus, the eEF1A-related ribosomal mechanisms were anticipated only by analogy with the bacterial homolog EF-Tu. Here, we report the first crystal structure of the mammalian eEF1A2*GDP complex which indicates major differences in the organization of the nucleotide-binding domain and intramolecular movements of eEF1A compared to EF-Tu. Our results explain the nucleotide exchange mechanism in the mammalian eEF1A and suggest that the first step of eEF1A*GDP dissociation from the 80S ribosome is the rotation of the nucleotide-binding domain observed after GTP hydrolysis.
Organizational Affiliation:
University of Grenoble Alpes, UVHCI, F-38000 Grenoble, France CNRS, UVHCI, F-38000 Grenoble, France Unit for Virus Host-Cell Interactions, University of Grenoble Alpes-EMBL-CNRS, 71 avenue des Martyrs, 38042 France.