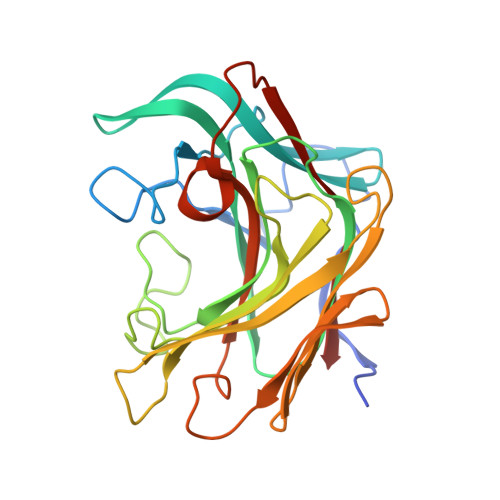
The Beta-Glucanase Zglama from Zobellia Galactanivorans Evolved a Bent Active Site Adapted for Efficient Degradation of Algal Laminarin
Labourel, A., Jam, M., Jeudy, A., Hehemann, J.H., Czjzek, M., Michel, G.(2014) J Biol Chem 289: 2027
- PubMed: 24337571
- DOI: https://doi.org/10.1074/jbc.M113.538843
- Primary Citation of Related Structures:
4BOW, 4BPZ, 4BQ1 - PubMed Abstract:
Laminarinase is commonly used to describe β-1,3-glucanases widespread throughout Archaea, bacteria, and several eukaryotic lineages. Some β-1,3-glucanases have already been structurally and biochemically characterized, but very few from organisms that are in contact with genuine laminarin, the storage polysaccharide of brown algae. Here we report the heterologous expression and subsequent biochemical and structural characterization of ZgLamAGH16 from Zobellia galactanivorans, the first GH16 laminarinase from a marine bacterium associated with seaweeds. ZgLamAGH16 contains a unique additional loop, compared with other GH16 laminarinases, which is composed of 17 amino acids and gives a bent shape to the active site cleft of the enzyme. This particular topology is perfectly adapted to the U-shaped conformation of laminarin chains in solution and thus explains the predominant specificity of ZgLamAGH16 for this substrate. The three-dimensional structure of the enzyme and two enzyme-substrate complexes, one with laminaritetraose and the other with a trisaccharide of 1,3-1,4-β-d-glucan, have been determined at 1.5, 1.35, and 1.13 Å resolution, respectively. The structural comparison of substrate recognition pattern between these complexes allows the proposition that ZgLamAGH16 likely diverged from an ancestral broad specificity GH16 β-glucanase and evolved toward a bent active site topology adapted to efficient degradation of algal laminarin.
Organizational Affiliation:
From Sorbonne Universités, UPMC Université Paris 06, UMR 7139, Marine Plants and Biomolecules, Station Biologique de Roscoff, F-29682 Roscoff, Bretagne, France and.