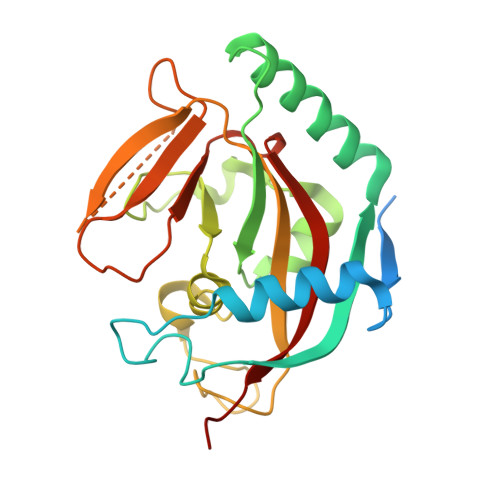
Evaluation and Structural Basis for the Inhibition of Tankyrases by Parp Inhibitors
Haikarainen, T., Narwal, M., Joensuu, P., Lehtio, L.(2014) ACS Med Chem Lett 5: 18
- PubMed: 24900770
- DOI: https://doi.org/10.1021/ml400292s
- Primary Citation of Related Structures:
4AVU, 4AVW, 4BJ9, 4BJB, 4BJC - PubMed Abstract:
Tankyrases, an enzyme subfamily of human poly(ADP-ribosyl)polymerases, are potential drug targets especially against cancer. We have evaluated inhibition of tankyrases by known PARP inhibitors and report five cocrystal structures of the most potent compounds in complex with human tankyrase 2. The inhibitors include the small general PARP inhibitors Phenanthridinone, PJ-34, and TIQ-A as well as the more advanced inhibitors EB-47 and rucaparib. The compounds anchor to the nicotinamide subsite of tankyrase 2. Crystal structures reveal flexibility of the ligand binding site with implications for drug development against tankyrases and other ADP-ribosyltransferases. EB-47 mimics the substrate NAD(+) and extends from the nicotinamide to the adenosine subsite. The clinical ARTD1 inhibitor candidate rucaparib was the most potent tankyrase inhibitor identified (24 and 14 nM for tankyrases), which indicates that inhibition of tankyrases would affect the cellular responses of this compound.
Organizational Affiliation:
Biocenter Oulu, Department of Biochemistry, University of Oulu , Oulu, Finland.