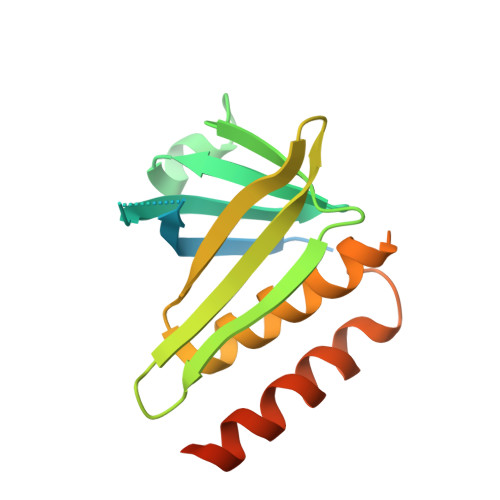
Structural and Functional Characterisation of the Kindlin-1 Pleckstrin Homology Domain
Yates, L.A., Lumb, C.N., Brahme, N.N., Zalyte, R., Bird, L.E., De Colibus, L., Owens, R.J., Calderwood, D.A., Sansom, M.S.P., Gilbert, R.J.C.(2012) J Biol Chem 287: 43246
- PubMed: 23132860
- DOI: https://doi.org/10.1074/jbc.M112.422089
- Primary Citation of Related Structures:
4BBK - PubMed Abstract:
Inside-out activation of integrins is mediated via the binding of talin and kindlin to integrin β-subunit cytoplasmic tails. The kindlin FERM domain is interrupted by a pleckstrin homology (PH) domain within its F2 subdomain. Here, we present data confirming the importance of the kindlin-1 PH domain for integrin activation and its x-ray crystal structure at a resolution of 2.1 Å revealing a C-terminal second α-helix integral to the domain but found only in the kindlin protein family. An isoform-specific salt bridge occludes the canonical phosphoinositide binding site, but molecular dynamics simulations display transient switching to an alternative open conformer. Molecular docking reveals that the opening of the pocket would enable potential ligands to bind within it. Although lipid overlay assays suggested the PH domain binds inositol monophosphates, surface plasmon resonance demonstrated weak affinities for inositol 3,4,5-triphosphate (Ins(3,4,5)P(3); K(D) ∼100 μM) and no monophosphate binding. Removing the salt bridge by site-directed mutagenesis increases the PH domain affinity for Ins(3,4,5)P(3) as measured by surface plasmon resonance and enables it to bind PtdIns(3,5)P(2) on a dot-blot. Structural comparison with other PH domains suggests that the phosphate binding pocket in the kindlin-1 PH domain is more occluded than in kindlins-2 and -3 due to its salt bridge. In addition, the apparent affinity for Ins(3,4,5)P(3) is affected by the presence of PO(4) ions in the buffer. We suggest the physiological ligand of the kindlin-1 PH domain is most likely not an inositol phosphate but another phosphorylated species.
Organizational Affiliation:
Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford OX3 7BN, United Kingdom.