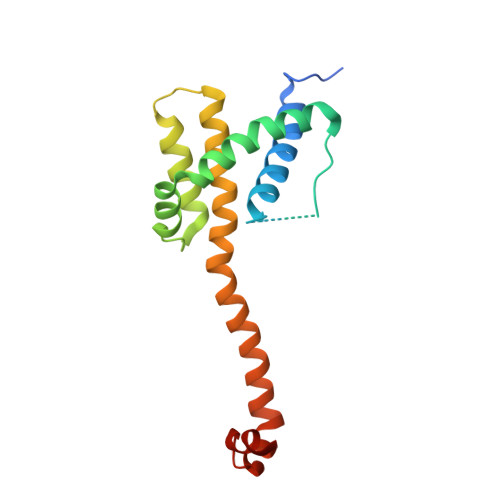
Crystal structure of Bax bound to the BH3 peptide of Bim identifies important contacts for interaction.
Robin, A.Y., Krishna Kumar, K., Westphal, D., Wardak, A.Z., Thompson, G.V., Dewson, G., Colman, P.M., Czabotar, P.E.(2015) Cell Death Dis 6: e1809-e1809
- PubMed: 26158515
- DOI: https://doi.org/10.1038/cddis.2015.141
- Primary Citation of Related Structures:
4ZIE, 4ZIF, 4ZIG, 4ZIH, 4ZII - PubMed Abstract:
The BH3-only protein Bim is a potent direct activator of the proapoptotic effector protein Bax, but the structural basis for its activity has remained poorly defined. Here we describe the crystal structure of the BimBH3 peptide bound to BaxΔC26 and structure-based mutagenesis studies. Similar to BidBH3, the BimBH3 peptide binds into the cognate surface groove of Bax using the conserved hydrophobic BH3 residues h1-h4. However, the structure and mutagenesis data show that Bim is less reliant compared with Bid on its 'h0' residues for activating Bax and that a single amino-acid difference between Bim and Bid encodes a fivefold difference in Bax-binding potency. Similar to the structures of BidBH3 and BaxBH3 bound to BaxΔC21, the structure of the BimBH3 complex with BaxΔC displays a cavity surrounded by Bax α1, α2, α5 and α8. Our results are consistent with a model in which binding of an activator BH3 domain to the Bax groove initiates separation of its core (α2-α5) and latch (α6-α8) domains, enabling its subsequent dimerisation and the permeabilisation of the mitochondrial outer membrane.
Organizational Affiliation:
1] The Walter and Eliza Hall Institute of Medical Research, Melbourne, VIC, Australia [2] Department of Medical Biology, The University of Melbourne, Melbourne, VIC, Australia.