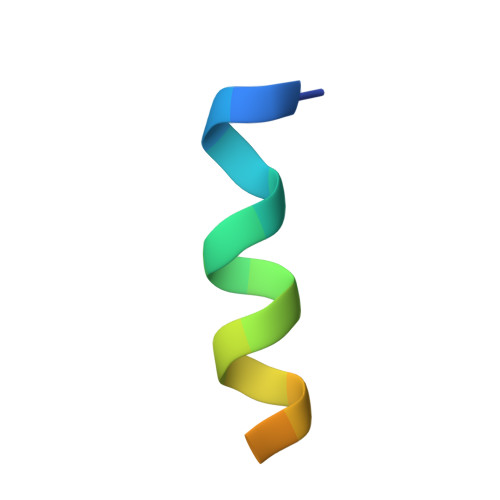
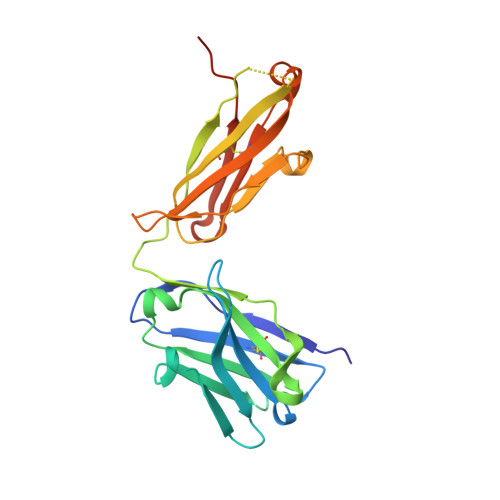
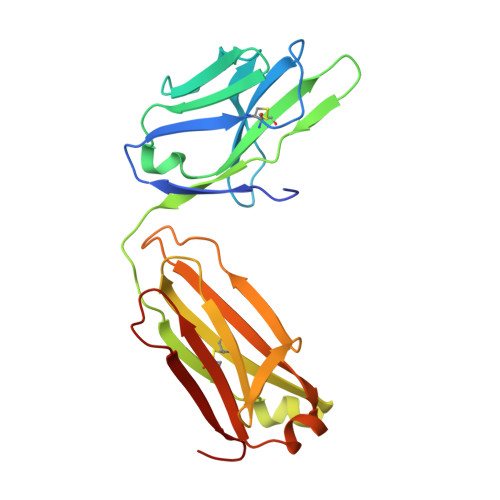
Engineering Synthetic Antibody Inhibitors Specific for LD2 or LD4 Motifs of Paxillin.
Nocula-Lugowska, M., Lugowski, M., Salgia, R., Kossiakoff, A.A.(2015) J Mol Biol 427: 2532-2547
- PubMed: 26087144
- DOI: https://doi.org/10.1016/j.jmb.2015.06.004
- Primary Citation of Related Structures:
4XGZ, 4XH2 - PubMed Abstract:
Focal adhesion protein paxillin links integrin and growth factor signaling to actin cytoskeleton. Most of paxillin signaling activity is regulated via leucine-rich LD motifs (LD1-LD5) located at the N-terminus. Here, we demonstrate a method to engineer highly selective synthetic antibodies (sABs) against LD2 and LD4 that are binding sites for focal adhesion kinase (FAK) and other proteins. Phage display selections against peptides were used to generate sABs recognizing each LD motif. In the obtained X-ray crystal structures of the LD-sAB complexes, the LD motifs are helical and bind sABs through a hydrophobic side, similarly as in the structures with natural paxillin partners. The sABs are capable of pulling down endogenous paxillin in complex with FAK and can visualize paxillin in focal adhesions in cells. They were also used as selective inhibitors to effectively compete with focal adhesion targeting domain of FAK for the binding to LD2 and LD4. The sABs are tools for investigation of paxillin LD binding "platforms" and are capable of inhibiting paxillin interactions, thereby useful as potential therapeutics in the future.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, The University of Chicago, Chicago, IL 60607, USA.