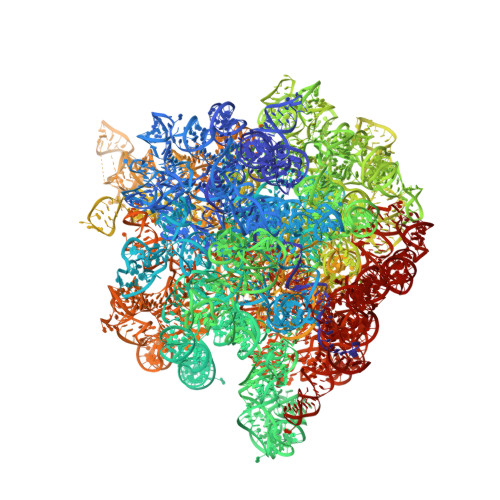
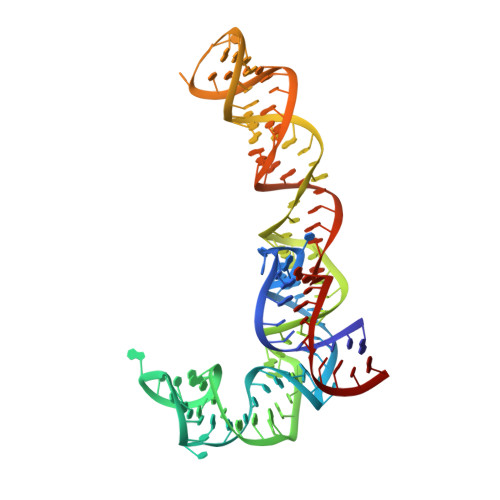
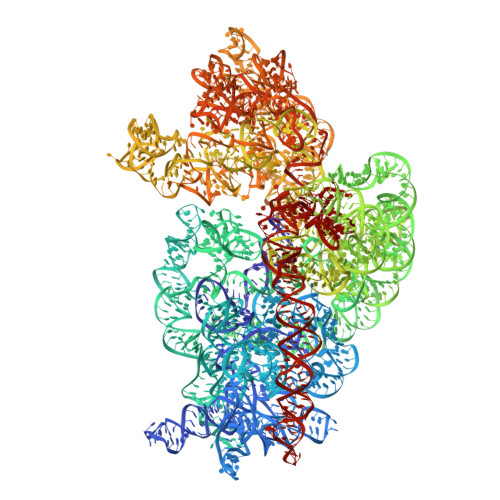
Binding and action of CEM-101, a new fluoroketolide antibiotic that inhibits protein synthesis.
Llano-Sotelo, B., Dunkle, J., Klepacki, D., Zhang, W., Fernandes, P., Cate, J.H., Mankin, A.S.(2010) Antimicrob Agents Chemother 54: 4961-4970
- PubMed: 20855725
- DOI: https://doi.org/10.1128/AAC.00860-10
- Primary Citation of Related Structures:
4WWW - PubMed Abstract:
We characterized the mechanism of action and the drug-binding site of a novel ketolide, CEM-101, which belongs to the latest class of macrolide antibiotics. CEM-101 shows high affinity for the ribosomes of Gram-negative (Escherichia coli) and Gram-positive (Staphylococcus aureus) bacteria. The ketolide shows high selectivity in its inhibitory action and readily interferes with synthesis of a reporter protein in the bacterial but not eukaryotic cell-free translation system. Binding of CEM-101 to its ribosomal target site was characterized biochemically and by X-ray crystallography. The X-ray structure of CEM-101 in complex with the E. coli ribosome shows that the drug binds in the major macrolide site in the upper part of the ribosomal exit tunnel. The lactone ring of the drug forms hydrophobic interactions with the walls of the tunnel, the desosamine sugar projects toward the peptidyl transferase center and interacts with the A2058/A2509 cleft, and the extended alkyl-aryl arm of the drug is oriented down the tunnel and makes contact with a base pair formed by A752 and U2609 of the 23S rRNA. The position of the CEM-101 alkyl-aryl extended arm differs from that reported for the side chain of the ketolide telithromycin complexed with either bacterial (Deinococcus radiodurans) or archaeal (Haloarcula marismortui) large ribosomal subunits but closely matches the position of the side chain of telithromycin complexed to the E. coli ribosome. A difference in the chemical structure of the side chain of CEM-101 in comparison with the side chain of telithromycin and the presence of the fluorine atom at position 2 of the lactone ring likely account for the superior activity of CEM-101. The results of chemical probing suggest that the orientation of the CEM-101 extended side chain observed in the E. coli ribosome closely resembles its placement in Staphylococcus aureus ribosomes and thus likely accurately reflects interaction of CEM-101 with the ribosomes of the pathogenic bacterial targets of the drug. Chemical probing further demonstrated weak binding of CEM-101, but not of erythromycin, to the ribosome dimethylated at A2058 by the action of Erm methyltransferase.
Organizational Affiliation:
Center for Pharmaceutical Biotechnology-m/c 870, University of Illinois, 900 S. Ashland Ave., Chicago, IL 60607, USA.