Crystal structure of C-terminal domain of penicillin binding protein Rv0907
Chang, C., Hatzos-Skintges, C., Jedrzejczak, R., Joachimiak, A., Midwest Center for Structural Genomics (MCSG)To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-lactamase | 102 | Mycobacterium tuberculosis H37Rv | Mutation(s): 0 Gene Names: P425_00947, RVBD_0907 | 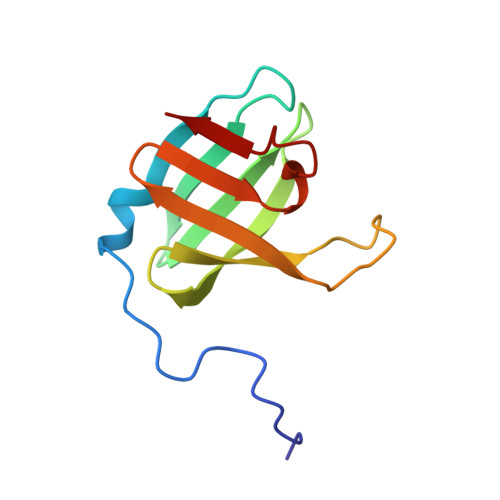 | |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BR Query on BR | B [auth A] C [auth A] D [auth A] E [auth A] F [auth A] | BROMIDE ION Br CPELXLSAUQHCOX-UHFFFAOYSA-M |  | ||
| NI Query on NI | R [auth A] | NICKEL (II) ION Ni VEQPNABPJHWNSG-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 35.305 | α = 90 |
| b = 49.839 | β = 90 |
| c = 59.345 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |