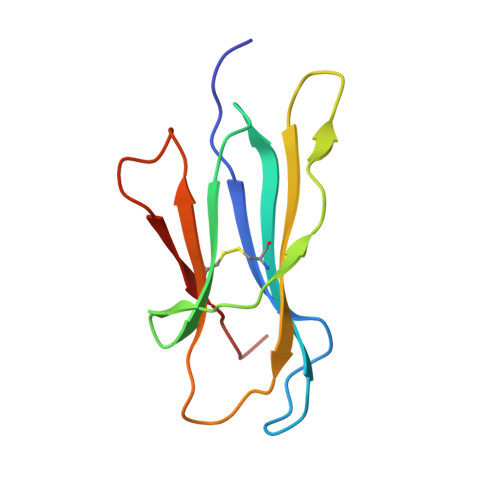
A covalent homodimer probing early oligomers along amyloid aggregation.
Halabelian, L., Relini, A., Barbiroli, A., Penco, A., Bolognesi, M., Ricagno, S.(2015) Sci Rep 5: 14651-14651
- PubMed: 26420657
- DOI: https://doi.org/10.1038/srep14651
- Primary Citation of Related Structures:
4R9H, 4RA3, 4RAH - PubMed Abstract:
Early oligomers are crucial in amyloid aggregation; however, due to their transient nature they are among the least structurally characterized species. We focused on the amyloidogenic protein beta2-microglobulin (β2m) whose early oligomers are still a matter of debate. An intermolecular interaction between D strands of facing β2m molecules was repeatedly observed, suggesting that such interface may be relevant for β2m dimerization. In this study, by mutating Ser33 to Cys, and assembling the disulphide-stabilized β2m homodimer (DimC33), such DD strand interface was locked. Although the isolated DimC33 display a stability similar to wt β2m under native conditions, it shows enhanced amyloid aggregation propensity. Three distinct crystal structures of DimC33 suggest that dimerization through the DD interface is instrumental for enhancing DimC33 aggregation propensity. Furthermore, the crystal structure of DimC33 in complex with the amyloid-specific dye Thioflavin-T pinpoints a second interface, which likely participates in the first steps of β2m aggregation. The present data provide new insight into β2m early steps of amyloid aggregation.
Organizational Affiliation:
Dipartimento di Bioscienze, Università degli Studi di Milano, Via Celoria 26, 20133 Milan, Italy.