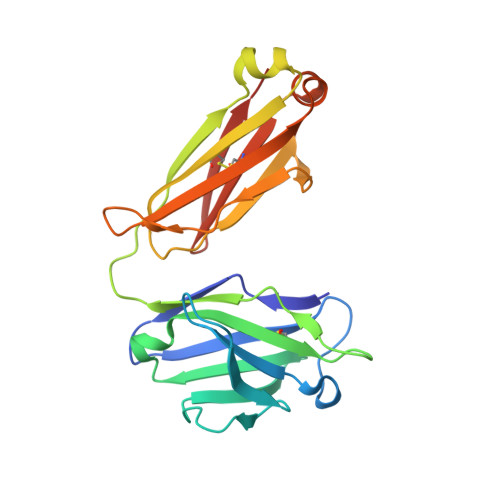
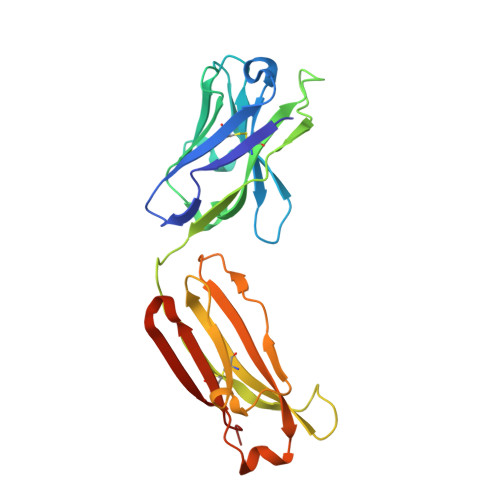
Camelid Ig V genes reveal significant human homology not seen in therapeutic target genes, providing for a powerful therapeutic antibody platform.
Klarenbeek, A., Mazouari, K.E., Desmyter, A., Blanchetot, C., Hultberg, A., de Jonge, N., Roovers, R.C., Cambillau, C., Spinelli, S., Del-Favero, J., Verrips, T., de Haard, H.J., Achour, I.(2015) MAbs 7: 693-706
- PubMed: 26018625
- DOI: https://doi.org/10.1080/19420862.2015.1046648
- Primary Citation of Related Structures:
4R90, 4R96 - PubMed Abstract:
Camelid immunoglobulin variable (IGV) regions were found homologous to their human counterparts; however, the germline V repertoires of camelid heavy and light chains are still incomplete and their therapeutic potential is only beginning to be appreciated. We therefore leveraged the publicly available HTG and WGS databases of Lama pacos and Camelus ferus to retrieve the germline repertoire of V genes using human IGV genes as reference. In addition, we amplified IGKV and IGLV genes to uncover the V germline repertoire of Lama glama and sequenced BAC clones covering part of the Lama pacos IGK and IGL loci. Our in silico analysis showed that camelid counterparts of all human IGKV and IGLV families and most IGHV families could be identified, based on canonical structure and sequence homology. Interestingly, this sequence homology seemed largely restricted to the Ig V genes and was far less apparent in other genes: 6 therapeutically relevant target genes differed significantly from their human orthologs. This contributed to efficient immunization of llamas with the human proteins CD70, MET, interleukin (IL)-1β and IL-6, resulting in large panels of functional antibodies. The in silico predicted human-homologous canonical folds of camelid-derived antibodies were confirmed by X-ray crystallography solving the structure of 2 selected camelid anti-CD70 and anti-MET antibodies. These antibodies showed identical fold combinations as found in the corresponding human germline V families, yielding binding site structures closely similar to those occurring in human antibodies. In conclusion, our results indicate that active immunization of camelids can be a powerful therapeutic antibody platform.
Organizational Affiliation:
a Department of Cell Biology; Utrecht University ; Utrecht , The Netherlands.