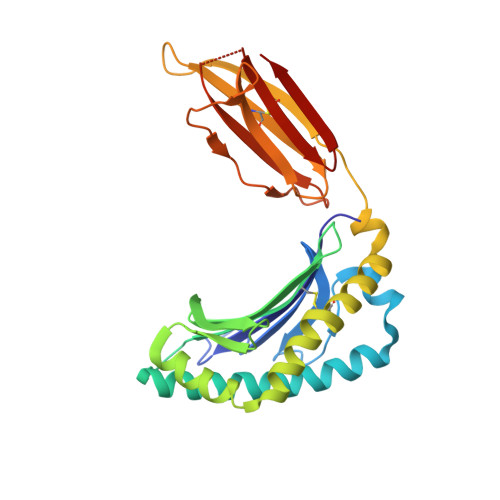
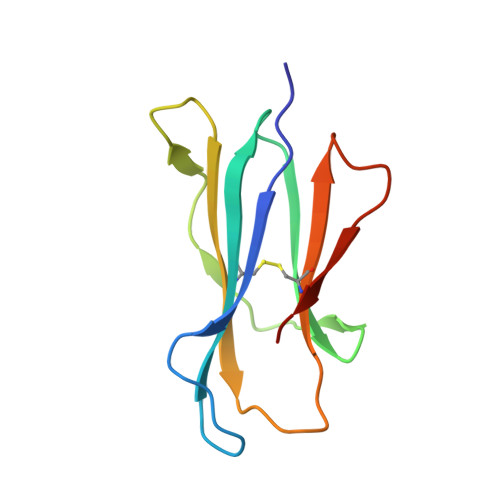
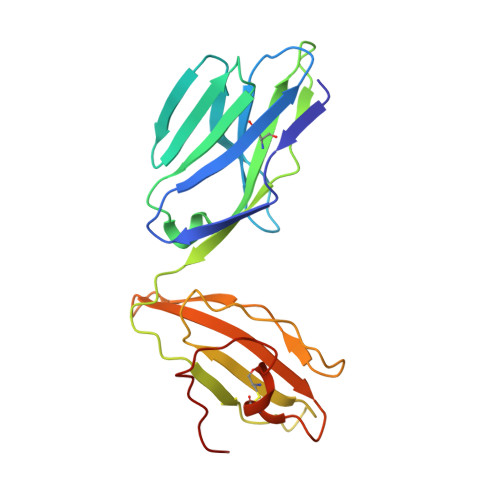
Antigen-loaded MR1 tetramers define T cell receptor heterogeneity in mucosal-associated invariant T cells.
Reantragoon, R., Corbett, A.J., Sakala, I.G., Gherardin, N.A., Furness, J.B., Chen, Z., Eckle, S.B., Uldrich, A.P., Birkinshaw, R.W., Patel, O., Kostenko, L., Meehan, B., Kedzierska, K., Liu, L., Fairlie, D.P., Hansen, T.H., Godfrey, D.I., Rossjohn, J., McCluskey, J., Kjer-Nielsen, L.(2013) J Exp Med 210: 2305-2320
- PubMed: 24101382
- DOI: https://doi.org/10.1084/jem.20130958
- Primary Citation of Related Structures:
4LCW - PubMed Abstract:
Mucosal-associated invariant T cells (MAIT cells) express a semi-invariant T cell receptor (TCR) α-chain, TRAV1-2-TRAJ33, and are activated by vitamin B metabolites bound by the major histocompatibility complex (MHC)-related class I-like molecule, MR1. Understanding MAIT cell biology has been restrained by the lack of reagents to specifically identify and characterize these cells. Furthermore, the use of surrogate markers may misrepresent the MAIT cell population. We show that modified human MR1 tetramers loaded with the potent MAIT cell ligand, reduced 6-hydroxymethyl-8-D-ribityllumazine (rRL-6-CH₂OH), specifically detect all human MAIT cells. Tetramer(+) MAIT subsets were predominantly CD8(+) or CD4(-)CD8(-), although a small subset of CD4(+) MAIT cells was also detected. Notably, most human CD8(+) MAIT cells were CD8α(+)CD8β(-/lo), implying predominant expression of CD8αα homodimers. Tetramer-sorted MAIT cells displayed a T(H)1 cytokine phenotype upon antigen-specific activation. Similarly, mouse MR1-rRL-6-CH₂OH tetramers detected CD4(+), CD4(-)CD8(-) and CD8(+) MAIT cells in Vα19 transgenic mice. Both human and mouse MAIT cells expressed a broad TCR-β repertoire, and although the majority of human MAIT cells expressed TRAV1-2-TRAJ33, some expressed TRAJ12 or TRAJ20 genes in conjunction with TRAV1-2. Accordingly, MR1 tetramers allow precise phenotypic characterization of human and mouse MAIT cells and revealed unanticipated TCR heterogeneity in this population.
Organizational Affiliation:
Department of Microbiology and Immunology, Peter Doherty Institute for Infection and Immunity; and 2 Department of Anatomy and Neuroscience; The University of Melbourne, Parkville, Victoria 3010, Australia.