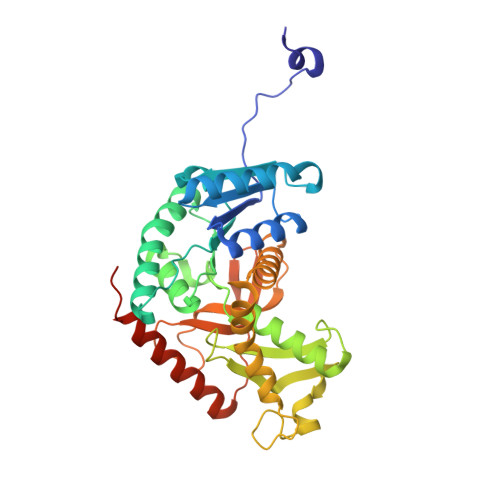
Structural characterization of the apo form and NADH binary complex of human lactate dehydrogenase.
Dempster, S., Harper, S., Moses, J.E., Dreveny, I.(2014) Acta Crystallogr D Biol Crystallogr 70: 1484-1490
- PubMed: 24816116
- DOI: https://doi.org/10.1107/S1399004714005422
- Primary Citation of Related Structures:
4L4R, 4L4S - PubMed Abstract:
Lactate dehydrogenase A (LDH-A) is a key enzyme in anaerobic respiration that is predominantly found in skeletal muscle and catalyses the reversible conversion of pyruvate to lactate in the presence of NADH. LDH-A is overexpressed in many tumours and has therefore emerged as an attractive target for anticancer drug discovery. Crystal structures of human LDH-A in the presence of inhibitors have been described, but currently no structures of the apo or binary NADH-bound forms are available for any mammalian LDH-A. Here, the apo structure of human LDH-A was solved at a resolution of 2.1 Å in space group P4122. The active-site loop adopts an open conformation and the packing and crystallization conditions suggest that the crystal form is suitable for soaking experiments. The soaking potential was assessed with the cofactor NADH, which yielded a ligand-bound crystal structure in the absence of any inhibitors. The structures show that NADH binding induces small conformational changes in the active-site loop and an adjacent helix. A comparison with other eukaryotic apo LDH structures reveals the conservation of intra-loop interactions. The structures provide novel insight into cofactor binding and provide the foundation for soaking experiments with fragments and inhibitors.
Organizational Affiliation:
School of Chemistry, University of Nottingham, Nottingham NG7 2RD, England.