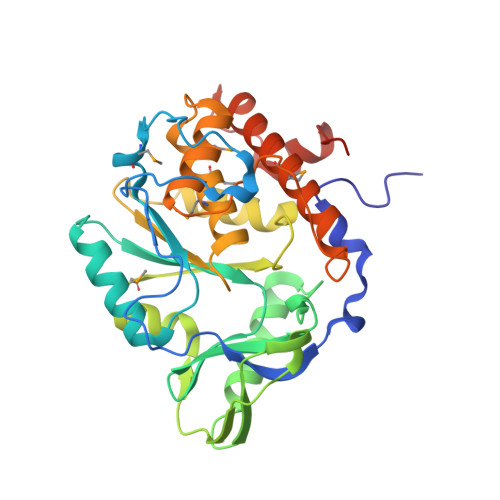
Crystal Structure of the computationally designed serine hydrolase 3mmj_2, Northeast Structural Genomics Consortium (NESG) Target OR318
Kuzin, A., Lew, S., Rajagopalan, S., Seetharaman, J., Maglaqui, M., Xiao, R., Lee, D., Everett, J.K., Acton, T.B., Montelione, G.T., Tong, L., Baker, D., Hunt, J.F.To be published.