Crystal and Solution Structures of Plasmodium falciparum Erythrocyte-binding Antigen 140 Reveal Determinants of Receptor Specificity during Erythrocyte Invasion.
Lin, D.H., Malpede, B.M., Batchelor, J.D., Tolia, N.H.(2012) J Biol Chem 287: 36830-36836
- PubMed: 22989878
- DOI: https://doi.org/10.1074/jbc.M112.409276
- Primary Citation of Related Structures:
4GF2 - PubMed Abstract:
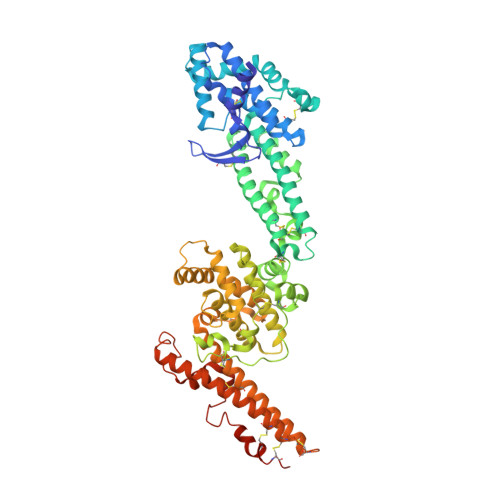
Erythrocyte-binding antigen 140 (PfEBA-140) is a critical Plasmodium falciparum erythrocyte invasion ligand that engages glycophorin C on host erythrocytes during malaria infection. The minimal receptor-binding region of PfEBA-140 contains two conserved Duffy binding-like (DBL) domains, a fold unique to Plasmodium species. Here, we present the crystal structure of the receptor-binding region of PfEBA-140 at 2.4 Å resolution. The two-domain binding region is present as a monomer in the asymmetric unit, and the structure reveals novel features in PfEBA-140 that are likely determinants of receptor specificity. Analysis by small-angle x-ray scattering demonstrated that the minimal binding region is monomeric in solution, consistent with the crystal structure. Erythrocyte binding assays showed that the full-length binding region containing the tandem DBL domains is required for erythrocyte engagement, suggesting that both domains contain critical receptor contact sites. The electrostatic surface of PfEBA-140 elucidates a basic patch that constitutes a putative high-affinity binding interface spanning both DBL domains. Mutation of residues within this interface results in severely diminished erythrocyte binding. This study provides insight into the structural basis and mechanism of PfEBA-140 receptor engagement and forms a basis for future studies of this critical interaction. In addition, the solution and crystal structures allow the first identification of likely determinants of erythrocyte receptor specificity for P. falciparum invasion ligands. A complete understanding of the PfEBA-140 erythrocyte invasion pathway will aid in the design of invasion inhibitory therapeutics and vaccines.
Organizational Affiliation:
Department of Molecular Microbiology, Washington University School of Medicine, St. Louis, Missouri 63110, USA.