Basis of Miscoding of the DNA Adduct N2,3-Ethenoguanine by Human Y-family DNA Polymerases.
Zhao, L., Pence, M.G., Christov, P.P., Wawrzak, Z., Choi, J.Y., Rizzo, C.J., Egli, M., Guengerich, F.P.(2012) J Biol Chem 287: 35516-35526
- PubMed: 22910910
- DOI: https://doi.org/10.1074/jbc.M112.403253
- Primary Citation of Related Structures:
4FS1, 4FS2 - PubMed Abstract:
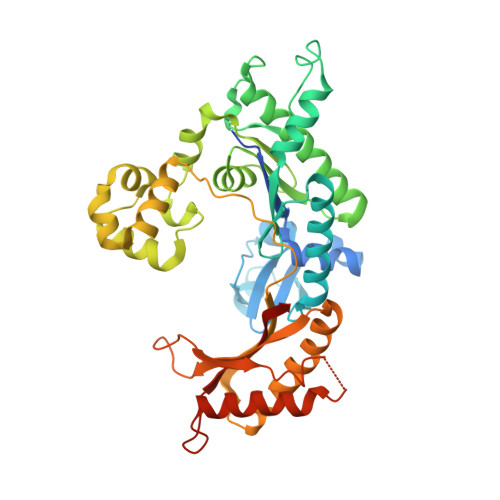
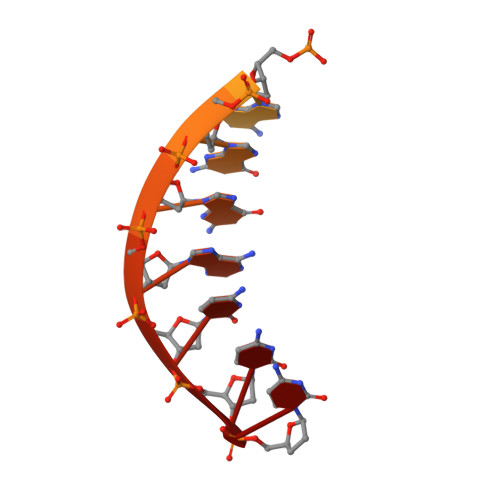
N(2),3-Ethenoguanine (N(2),3-εG) is one of the exocyclic DNA adducts produced by endogenous processes (e.g. lipid peroxidation) and exposure to bioactivated vinyl monomers such as vinyl chloride, which is a known human carcinogen. Existing studies exploring the miscoding potential of this lesion are quite indirect because of the lability of the glycosidic bond. We utilized a 2'-fluoro isostere approach to stabilize this lesion and synthesized oligonucleotides containing 2'-fluoro-N(2),3-ε-2'-deoxyarabinoguanosine to investigate the miscoding potential of N(2),3-εG by Y-family human DNA polymerases (pols). In primer extension assays, pol η and pol κ replicated through N(2),3-εG, whereas pol ι and REV1 yielded only 1-base incorporation. Steady-state kinetics revealed that dCTP incorporation is preferred opposite N(2),3-εG with relative efficiencies in the order of pol κ > REV1 > pol η ≈ pol ι, and dTTP misincorporation is the major miscoding event by all four Y-family human DNA pols. Pol ι had the highest dTTP misincorporation frequency (0.71) followed by pol η (0.63). REV1 misincorporated dTTP and dGTP with much lower frequencies. Crystal structures of pol ι with N(2),3-εG paired to dCTP and dTTP revealed Hoogsteen-like base pairing mechanisms. Two hydrogen bonds were observed in the N(2),3-εG:dCTP base pair, whereas only one appears to be present in the case of the N(2),3-εG:dTTP pair. Base pairing mechanisms derived from the crystal structures explain the slightly favored dCTP insertion for pol ι in steady-state kinetic analysis. Taken together, these results provide a basis for the mutagenic potential of N(2),3-εG.
Organizational Affiliation:
Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee 37232-0146; Department of Center in Molecular Toxicology, Vanderbilt University School of Medicine, Nashville, Tennessee 37232-0146.