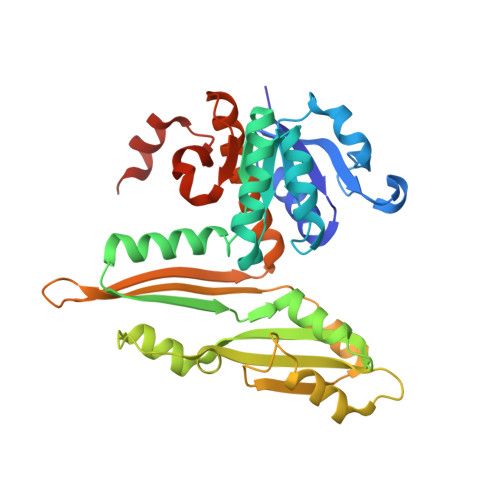
Structural insight into the thermostable NADP(+)-dependent meso-diaminopimelate dehydrogenase from Ureibacillus thermosphaericus
Akita, H., Seto, T., Ohshima, T., Sakuraba, H.(2015) Acta Crystallogr D Biol Crystallogr 71: 1136-1146
- PubMed: 25945579
- DOI: https://doi.org/10.1107/S1399004715003673
- Primary Citation of Related Structures:
3WYB, 3WYC - PubMed Abstract:
Crystal structures of the thermostable meso-diaminopimelate dehydrogenase (DAPDH) from Ureibacillus thermosphaericus were determined for the enzyme in the apo form and in complex with NADP(+) and N-tris(hydroxymethyl)methyl-2-aminoethanesulfonic acid. The main-chain coordinates of the enzyme showed notable similarity to those of Symbiobacterium thermophilum DAPDH. However, the subunit arrangement of U. thermosphaericus DAPDH (a dimer) was totally different from that of the S. thermophilum enzyme (a hexamer). Structural comparison with the dimeric enzyme from the mesophile Corynebacterium glutamicum revealed that the presence of large numbers of intrasubunit and intersubunit hydrophobic interactions, as well as the extensive formation of intersubunit ion-pair networks, were likely to be the main factors contributing to the higher thermostability of U. thermosphaericus DAPDH. This differs from S. thermophilum DAPDH, within which the unique hexameric assembly is likely to be responsible for its high thermostability. Analysis of the active site of U. thermosphaericus DAPDH revealed the key factors responsible for the marked difference in substrate specificity between DAPDH and the D-amino acid dehydrogenase recently created from DAPDH by introducing five point mutations [Akita et al. (2012). Biotechnol. Lett. 34, 1693-1699; 1701-1702].
Organizational Affiliation:
Applied Molecular Microbiology and Biomass Chemistry, Bioscience and Biotechnology, Faculty of Agriculture, Kyushu University, 6-10-1 Hakozaki, Higashi-ku, Fukuoka 812-8581, Japan.