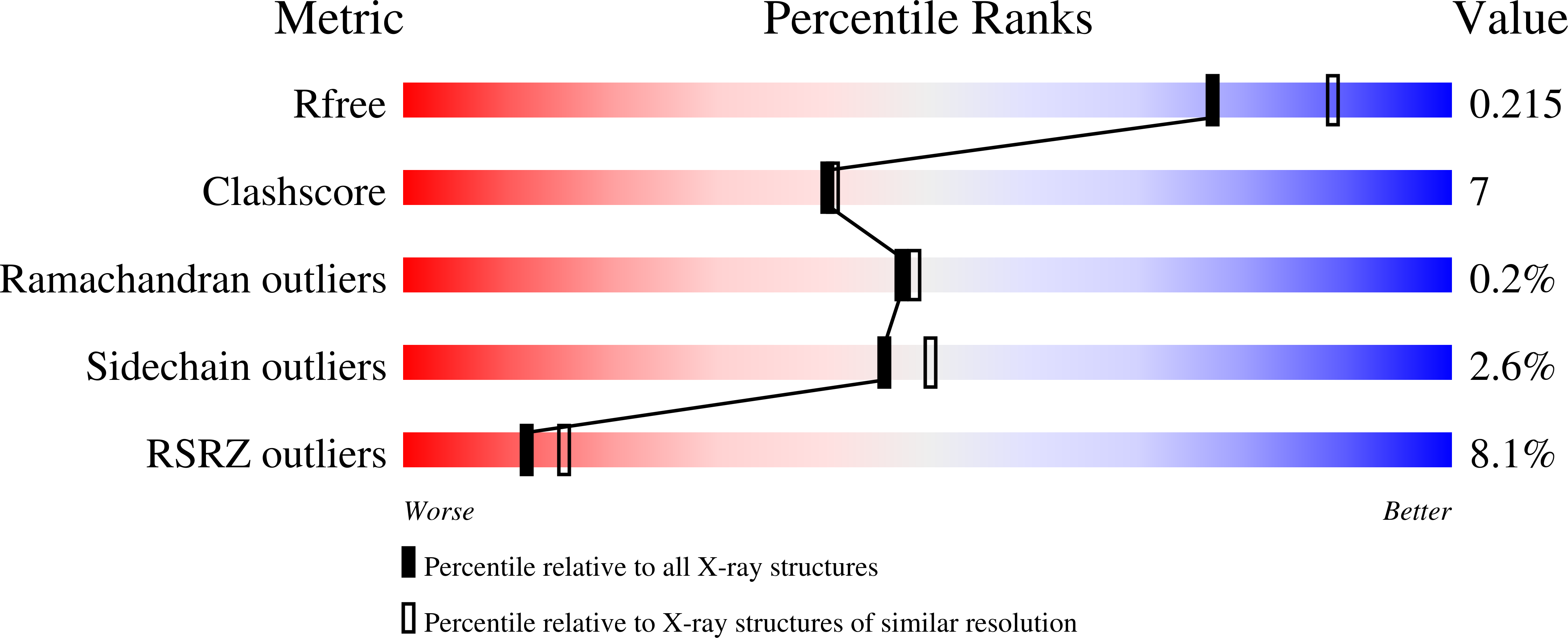
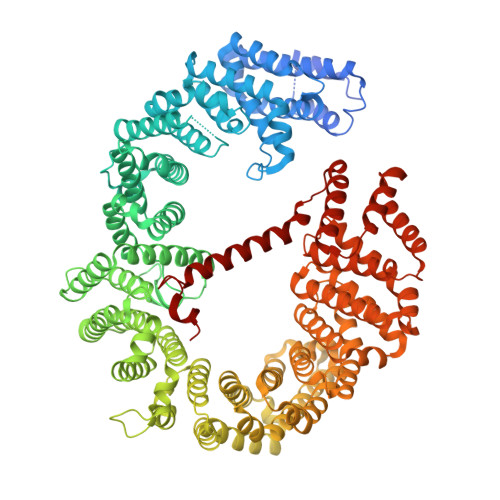
A 2.1- angstrom -resolution crystal structure of unliganded CRM1 reveals the mechanism of autoinhibition
Saito, N., Matsuura, Y.(2013) J Mol Biol 425: 350-364
- PubMed: 23164569
- DOI: https://doi.org/10.1016/j.jmb.2012.11.014
- Primary Citation of Related Structures:
3VYC - PubMed Abstract:
CRM1 mediates nuclear export of numerous proteins and ribonucleoproteins containing a leucine-rich nuclear export signal (NES). Binding of RanGTP to CRM1 in the nucleus stabilizes cargo association with CRM1, and vice versa, but the mechanism underlying the positive cooperativity in RanGTP and NES binding to CRM1 remains incompletely understood. Herein we report a 2.1-Å-resolution crystal structure of unliganded Saccharomyces cerevisiae CRM1 (Xpo1p) that demonstrates that an internal loop of CRM1 (referred to as HEAT9 loop) is primarily responsible for maintaining the NES-binding cleft in a closed conformation, rendering CRM1 incapable of NES binding in the absence of RanGTP. The structure also shows that the C-terminal tail of CRM1 stabilizes the autoinhibitory conformation of the HEAT9 loop and thereby reinforces autoinhibition. Comparison with the structures of CRM1-NES-RanGTP complexes reveals how binding of RanGTP is associated with a series of allosteric conformational changes in CRM1 that lead to opening of the NES-binding cleft, allowing for stable binding of NES cargoes.
Organizational Affiliation:
Division of Biological Science, Graduate School of Science, Nagoya University, Furo-cho, Chikusa-ku, Nagoya 464-8602, Japan.