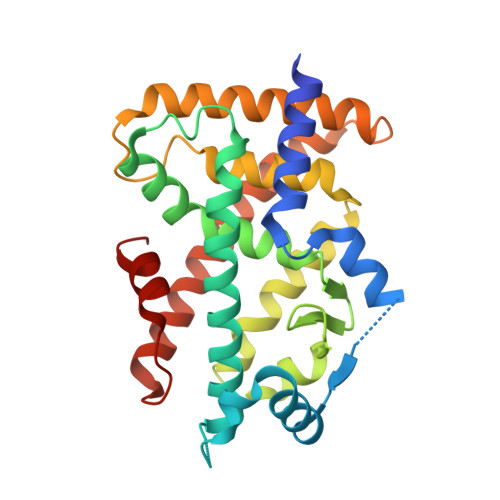
Peroxisome proliferator-activated receptors (PPARs) have multiple binding points that accommodate ligands in various conformations: phenylpropanoic acid-type PPAR ligands bind to PPAR in different conformations, depending on the subtype
Kuwabara, N., Oyama, T., Tomioka, D., Ohashi, M., Yanagisawa, J., Shimizu, T., Miyachi, H.(2012) J Med Chem 55: 893-902
- PubMed: 22185225
- DOI: https://doi.org/10.1021/jm2014293
- Primary Citation of Related Structures:
3VI8, 3VJH, 3VJI - PubMed Abstract:
Human peroxisome proliferator-activated receptors (hPPARs) are ligand-dependent transcription factors that control various biological responses, and there are three subtypes: hPPARα, hPPARδ, and hPPARγ. We report here that α-substituted phenylpropanoic acid-type hPPAR agonists with similar structure bind to the hPPAR ligand binding domain (LBD) in different conformations, depending on the receptor subtype. These results might indicate that hPPAR ligand binding pockets have multiple binding points that can be utilized to accommodate structurally flexible hPPAR ligands.
Organizational Affiliation:
Graduate School of Pharmaceutical Sciences, Faculty of Pharmaceutical Sciences, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.