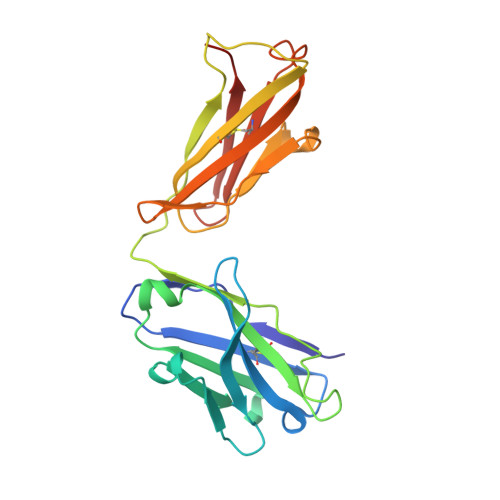
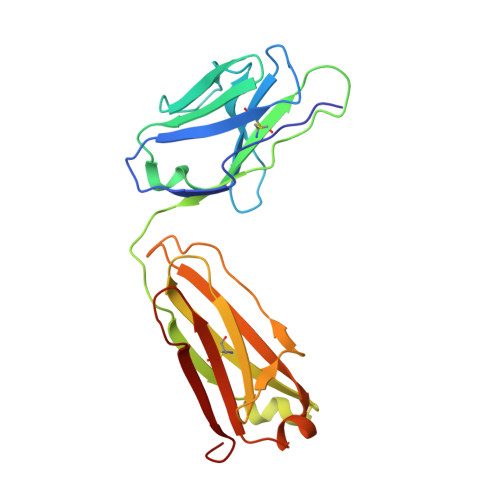
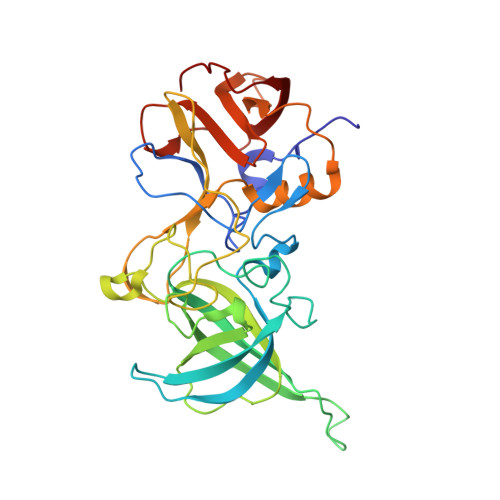
Structural Basis for Broad Detection of Genogroup II Noroviruses by a Monoclonal Antibody That Binds to a Site Occluded in the Viral Particle.
Hansman, G.S., Taylor, D.W., McLellan, J.S., Smith, T.J., Georgiev, I., Tame, J.R., Park, S.Y., Yamazaki, M., Gondaira, F., Miki, M., Katayama, K., Murata, K., Kwong, P.D.(2012) J Virol 86: 3635-3646
- PubMed: 22278249
- DOI: https://doi.org/10.1128/JVI.06868-11
- Primary Citation of Related Structures:
3V7A - PubMed Abstract:
Human noroviruses are genetically and antigenically highly divergent. Monoclonal antibodies raised in mice against one kind of norovirus virus-like particle (VLP), however, were found to have broad recognition. In this study, we present the crystal structure of the antigen-binding fragment (Fab) for one of these broadly reactive monoclonal antibodies, 5B18, in complex with the capsid-protruding domain from a genogroup II genotype 10 (GII.10) norovirus at 3.3-Å resolution and, also, the cryo-electron microscopy structure of the GII.10 VLP at ∼10-Å resolution. The GII.10 VLP structure was more similar in overall architecture to the GV.1 murine norovirus virion than to the prototype GI.1 human norovirus VLP, with the GII.10 protruding domain raised ∼15 Å off the shell domain and rotated ∼40° relative to the GI.1 protruding domain. In the crystal structure, the 5B18 Fab bound to a highly conserved region of the protruding domain. Based on the VLP structure, this region is involved in interactions with other regions of the capsid and is buried in the virus particle. Despite the occluded nature of the recognized epitope in the VLP structure, enzyme-linked immunosorbent assay (ELISA) binding suggested that the 5B18 antibody was able to capture intact VLPs. Together, the results provide evidence that the norovirus particle is capable of extreme conformational flexibility, which may allow for antibody recognition of conserved surfaces that would otherwise be buried on intact particles.
Organizational Affiliation:
Department of Virology II, National Institute of Infectious Diseases, Tokyo, Japana.