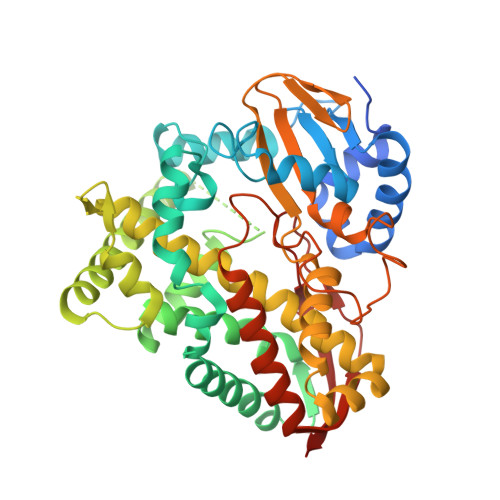
Structure and function of CYP108D1 from Novosphingobium aromaticivorans DSM12444: an aromatic hydrocarbon-binding P450 enzyme
Bell, S.G., Yang, W., Yorke, J.A., Zhou, W., Wang, H., Harmer, J., Copley, R., Zhang, A., Zhou, R., Bartlam, M., Rao, Z., Wong, L.-L.(2012) Acta Crystallogr D Biol Crystallogr 68: 277-291
- PubMed: 22349230
- DOI: https://doi.org/10.1107/S090744491200145X
- Primary Citation of Related Structures:
3TKT - PubMed Abstract:
CYP108D1 from Novosphingobium aromaticivorans DSM12444 binds a range of aromatic hydrocarbons such as phenanthrene, biphenyl and phenylcyclohexane. Its structure, which is reported here at 2.2 Å resolution, is closely related to that of CYP108A1 (P450terp), an α-terpineol-oxidizing enzyme. The compositions and structures of the active sites of these two enzymes are very similar; the most significant changes are the replacement of Glu77 and Thr103 in CYP108A1 by Thr79 and Val105 in CYP108D1. Other residue differences lead to a larger and more hydrophobic access channel in CYP108D1. These structural features are likely to account for the weaker α-terpineol binding by CYP108D1 and, when combined with the presence of three hydrophobic phenylalanine residues in the active site, promote the binding of aromatic hydrocarbons. The haem-proximal surface of CYP108D1 shows a different charge distribution and topology to those of CYP101D1, CYP101A1 and CYP108A1, including a pronounced kink in the proximal loop of CYP108D1, which may result in poor complementarity with the [2Fe-2S] ferredoxins Arx, putidaredoxin and terpredoxin that are the respective redox partners of these three P450 enzymes. The unexpectedly low reduction potential of phenylcyclohexane-bound CYP108D1 (-401 mV) may also contribute to the low activity observed with these ferredoxins. CYP108D1 appears to function as an aromatic hydrocarbon hydroxylase that requires a different electron-transfer cofactor protein.
Organizational Affiliation:
Inorganic Chemistry Laboratory, Department of Chemistry, University of Oxford,South Parks Road, Oxford OX1 3QR, England. stephen.bell@chem.ox.ac.uk