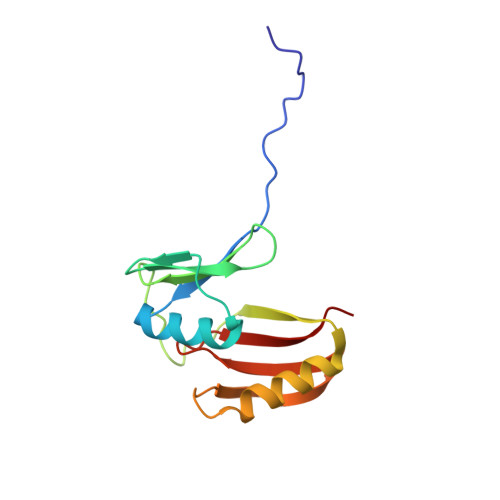
Atomic resolution structure of EhpR: phenazine resistance in Enterobacter agglomerans Eh1087 follows principles of bleomycin / mitomycin C resistance in other bacteria.
Yu, S., Vit, A., Devenish, S., Mahanty, H.K., Itzen, A., Goody, R.S., Blankenfeldt, W.(2011) BMC Struct Biol 11: 33-33
- PubMed: 21849072
- DOI: https://doi.org/10.1186/1472-6807-11-33
- Primary Citation of Related Structures:
3SK1, 3SK2 - PubMed Abstract:
The phenazines are redox-active secondary metabolites that a large number of bacterial strains produce and excrete into the environment. They possess antibiotic activity owing to the fact that they can reduce molecular oxygen to toxic reactive oxygen species. In order to take advantage of this activity, phenazine producers need to protect themselves against phenazine toxicity. Whereas it is believed that phenazine-producing pseudomonads possess highly active superoxide dismutases and catalases, it has recently been found that the plant-colonizing bacterium Enterobacter agglomerans expresses a small gene ehpR to render itself resistant towards D-alanyl-griseoluteic acid, the phenazine antibiotic produced by this strain.
Organizational Affiliation:
Department of Physical Biochemistry, Max Planck Institute of Molecular Physiology, Otto-Hahn-Straße 11, 44227 Dortmund, Germany.