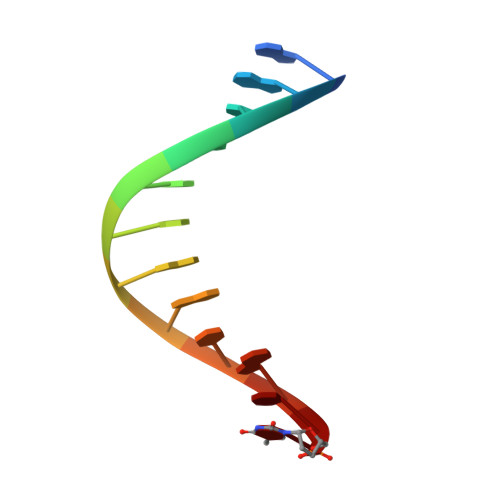
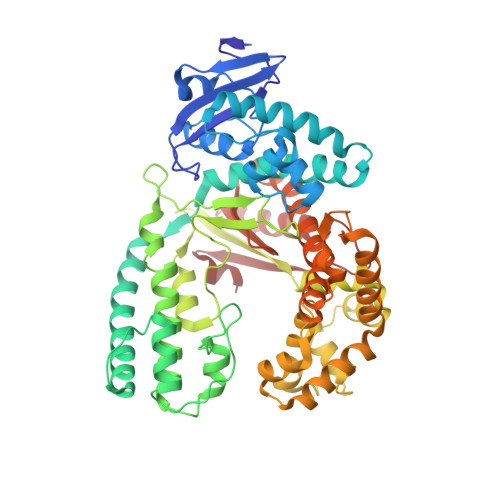
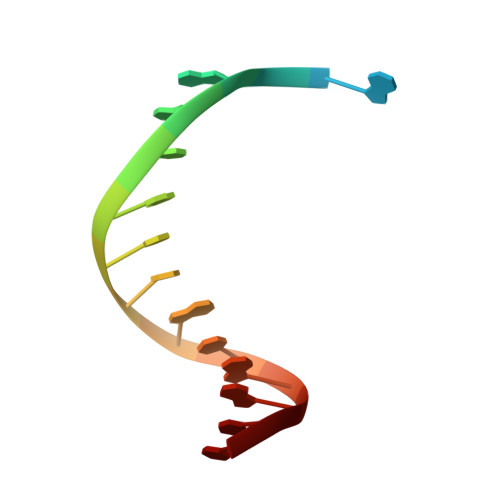
Amino Acid templating mechanisms in selection of nucleotides opposite abasic sites by a family a DNA polymerase.
Obeid, S., Welte, W., Diederichs, K., Marx, A.(2012) J Biol Chem 287: 14099-14108
- PubMed: 22318723
- DOI: https://doi.org/10.1074/jbc.M111.334904
- Primary Citation of Related Structures:
3RR7, 3RR8, 3RRG, 3RRH, 3T3F - PubMed Abstract:
Cleavage of the N-glycosidic bond that connects the nucleobase to the backbone in DNA leads to abasic sites, the most frequent lesion under physiological conditions. Several DNA polymerases preferentially incorporate an A opposite this lesion, a phenomenon termed "A-rule." Accordingly, KlenTaq, the large fragment of Thermus aquaticus DNA polymerase I, incorporates a nucleotide opposite an abasic site with efficiencies of A > G > T > C. Here we provide structural insights into constraints of the active site during nucleotide selection opposite an abasic site. It appears that these confines govern the nucleotide selection mainly by interaction of the incoming nucleotide with Tyr-671. Depending on the nucleobase, the nucleotides are differently positioned opposite Tyr-671 resulting in different alignments of the functional groups that are required for bond formation. The distances between the α-phosphate and the 3'-primer terminus increases in the order A < G < T, which follows the order of incorporation efficiency. Additionally, a binary KlenTaq structure bound to DNA containing an abasic site indicates that binding of the nucleotide triggers a remarkable rearrangement of enzyme and DNA template. The ability to resolve the stacking arrangement might be dependent on the intrinsic properties of the respective nucleotide contributing to nucleotide selection. Furthermore, we studied the incorporation of a non-natural nucleotide opposite an abasic site. The nucleotide was often used in studying stacking effects in DNA polymerization. Here, no interaction with Tyr-761 as found for the natural nucleotides is observed, indicating a different reaction path for this non-natural nucleotide.
Organizational Affiliation:
Department of Chemistry, University of Konstanz, Universita¨tsstrasse 10, D 78457 Konstanz, Germany.