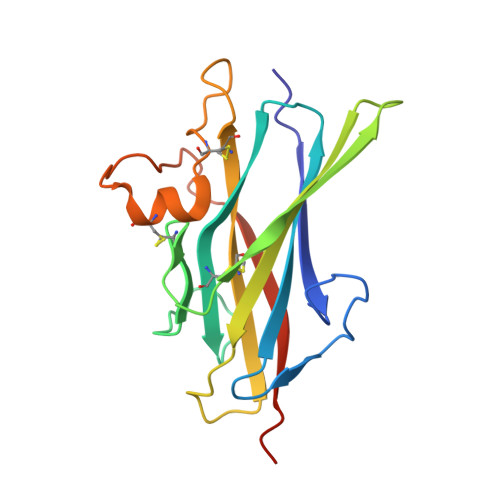
Structure of betaglycan zona pellucida (ZP)-C domain provides insights into ZP-mediated protein polymerization and TGF-{beta} binding.
Lin, S.J., Hu, Y., Zhu, J., Woodruff, T.K., Jardetzky, T.S.(2011) Proc Natl Acad Sci U S A 108: 5232-5236
- PubMed: 21402931
- DOI: https://doi.org/10.1073/pnas.1010689108
- Primary Citation of Related Structures:
3QW9 - PubMed Abstract:
The zona pellucida (ZP) domain is a bipartite protein structural element comprised of ZP-N and ZP-C regions. Most notable for its ability to mediate protein polymerization, many ZP proteins polymerize and assemble into long fibrils that form specialized extracellular matrices. Other ZP proteins (namely, betaglycan and endoglin) do not polymerize but serve as important membrane coreceptors for ligands in the transforming growth factor-β (TGF-β) superfamily. Here, we present the 2.0-Å resolution crystal structure of the betaglycan ZP-C region in combination with a downstream region known as the external hydrophobic patch (EHP). Similar to the ZP-N region, the ZP-C region also adopts an immunoglobulin-like fold, despite sharing no sequence homology and possessing different disulfide linkages. The EHP region, which was previously thought to be external to the ZP region, is integral to the ZP-C domain and corresponds to the ZP-C G strand. Our structure also indicates that the critical maturation cleavage of ZP proteins, a process that activates nascent ZP proteins for polymerization, occurs within the immunoglobulin domain at the FG loop. Nonpolymerizing ZP proteins such as betaglycan and endoglin do not contain this cleavage site. Finally, our structure suggests that the AB loop and the convex surface pocket are regions important for TGF-β ligand binding.
Organizational Affiliation:
Department of Structural Biology, Stanford University School of Medicine, 371 Serra Mall, Stanford, CA 94305, USA.